Practice: Chp. 15#
The answers to the questions in this chapter were originally developed with rstan, but running all
the cells at once would consistently produce a crash (greater than half the time). The error was
similar to:
*** caught segfault ***
address 0x7fd4069fa9d0, cause 'memory not mapped'
An irrecoverable exception occurred. R is aborting now ...
Stan developers advise moving to CmdStan to avoid these crashes:
Installation instructions for cmdstan and cmdstanr:
This environment includes cmdstan in conda:
library(cmdstanr)
set_cmdstan_path("/opt/conda/bin/cmdstan")
This is cmdstanr version 0.4.0
- Online documentation and vignettes at mc-stan.org/cmdstanr
- Use set_cmdstan_path() to set the path to CmdStan
- Use install_cmdstan() to install CmdStan
CmdStan path set to: /opt/conda/bin/cmdstan
After switching to CmdStan, the crashes completely went away. For more details on the process, see the commits near this change. The warning messages you see in this chapter may look otherwise unfamiliar.
source("iplot.R")
suppressPackageStartupMessages(library(rethinking))
15E1. Rewrite the Oceanic tools model (from Chapter 11) below so that it assumes measured error on the log population sizes of each society. You don’t need to fit the model to data. Just modify the mathematical formula below.
Answer. Assuming a \(P_{SE,i}\) is available, though it is not in the data:
15E2. Rewrite the same model so that it allows imputation of missing values for log population. There aren’t any missing values in the variable, but you can still write down a model formula that would imply imputation, if any values were missing.
Answer. As a challenge we’ll add this to model that also considers measurement error, assuming either both the observed value and the measurement are present, or neither. The priors could be much better:
15M1. Using the mathematical form of the imputation model in the chapter, explain what is being assumed about how the missing values were generated.
ERROR. It’s not clear what the author means by ‘the imputation model’; there are several models in the text.
Answer. In section 15.2.2. the first imputation model assumes a normal distribution for brain sizes. This is somewhat unreasonable, as explained in the text, because these values should be bounded between zero and one.
In the second model for this primate example we still assume a normal distribution, but now correlated with body mass.
15M2. Reconsider the primate milk missing data example from the chapter. This time, assign \(B\) a distribution that is properly bounded between zero and 1. A beta distribution, for example, is a good choice.
Answer. This question requires a start= list to run at all, which isn’t suggested for these
imputation models until a later question. We use the dbeta2 parameterization provided by ulam to
make the parameters a bit more understandable (as opposed to dbeta). Fitting the model:
data(milk)
d <- milk
d$neocortex.prop <- d$neocortex.perc / 100
d$logmass <- log(d$mass)
stdB = standardize(d$neocortex.prop)
dat_list <- list(
K = standardize(d$kcal.per.g),
B = d$neocortex.prop,
M = standardize(d$logmass),
center = attr(stdB, "scaled:center"),
scale = attr(stdB, "scaled:scale")
)
m_milk_beta <- ulam(
alist(
K ~ dnorm(mu, sigma),
mu <- a + bB*B + bM*M,
B ~ dbeta2(p, theta),
a ~ dnorm(0, 0.5),
c(bB, bM) ~ dnorm(0, 0.5),
p ~ dunif(0, 1),
theta ~ dunif(0, 1000),
sigma ~ dexp(1)
), cmdstan=TRUE, data=dat_list, chains=4, cores=4, start=list(B_impute=rep(0.7,12))
)
Found 12 NA values in B and attempting imputation.
Without bounds on the parameters to the beta distribution, this model produces a significant number
of divergent transitions. To avoid these errors, we are using uniform priors on the two parameters
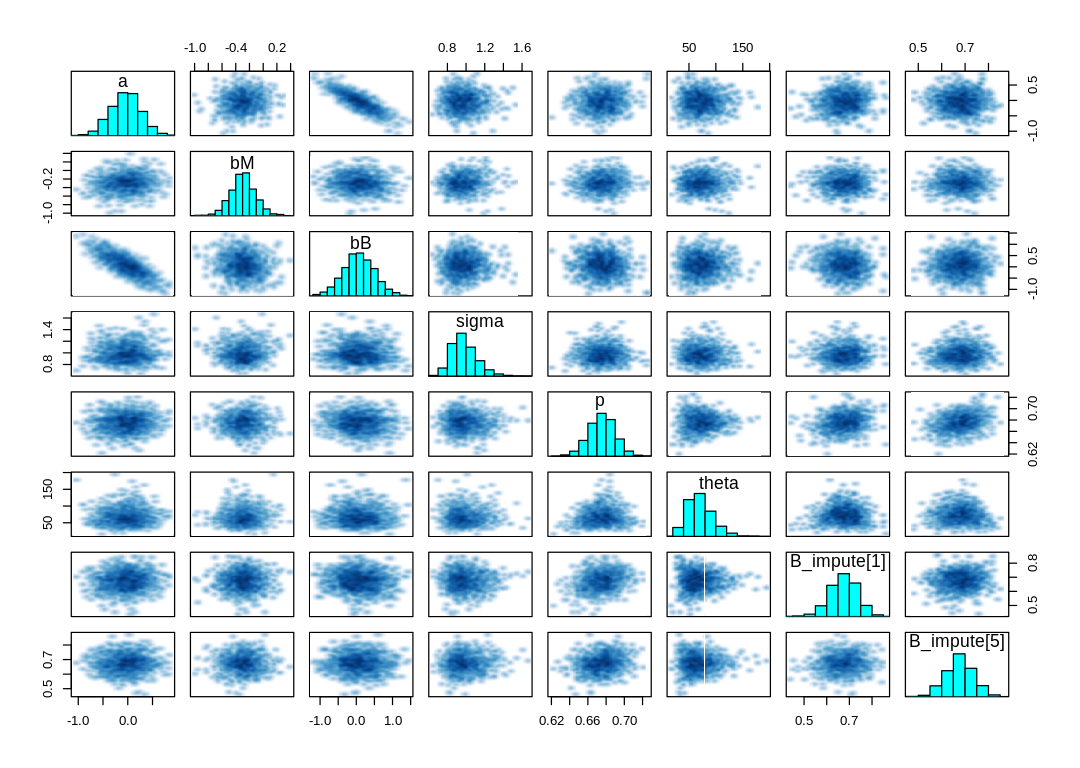
to the beta distribution. Checking the pairs and traceplot:
sel_pars=c("a", "bM", "bB", "sigma", "p", "theta", "B_impute[1]", "B_impute[5]")
iplot(function() {
pairs(m_milk_beta@stanfit, pars=sel_pars)
})
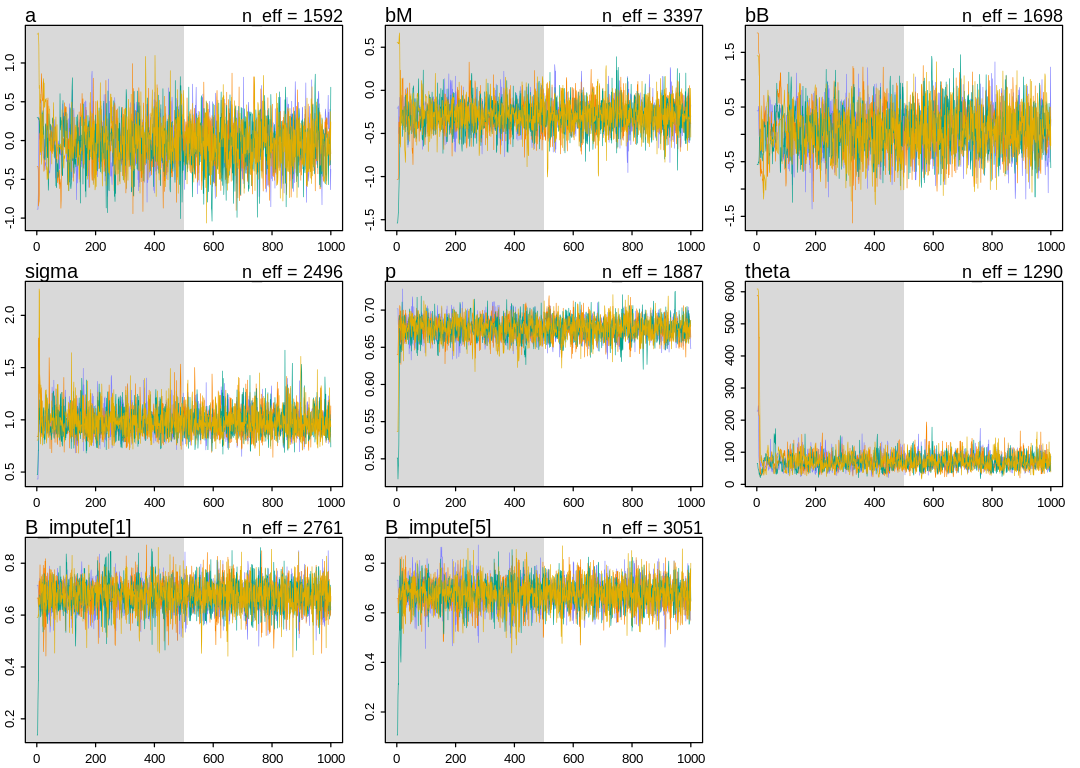
iplot(function() {
traceplot(m_milk_beta, pars=sel_pars)
})
[1] 1000
[1] 1
[1] 1000
The precis results:
display(precis(m_milk_beta, depth=3), mimetypes="text/plain")
iplot(function() {
plot(precis(m_milk_beta, depth=3), main="m_milk_beta")
}, ar=2)
mean sd 5.5% 94.5% n_eff
a -0.04295665 0.30361899 -0.5376136 0.426910758 1592.433
bM -0.29418840 0.18302593 -0.5798064 -0.002633437 3397.419
bB 0.07365296 0.43005402 -0.6299967 0.762377178 1698.291
p 0.67533351 0.01442545 0.6522330 0.697699699 1886.658
theta 70.77433602 23.50242292 38.0426798 111.287303705 1289.928
sigma 0.98056955 0.13683458 0.7918979 1.216318996 2495.705
B_impute[1] 0.67454671 0.06042883 0.5781118 0.766966686 2760.941
B_impute[2] 0.67520062 0.05694163 0.5842362 0.764997764 3034.462
B_impute[3] 0.67567915 0.05998694 0.5766410 0.770490172 3037.513
B_impute[4] 0.67604852 0.05811344 0.5790328 0.765995901 3214.573
B_impute[5] 0.67626441 0.05852606 0.5807795 0.768901601 3051.055
B_impute[6] 0.67604027 0.06213170 0.5747488 0.767945883 2282.398
B_impute[7] 0.67492820 0.05825896 0.5816310 0.766487154 3311.031
B_impute[8] 0.67587230 0.05888821 0.5780833 0.765306819 3140.502
B_impute[9] 0.67528655 0.05869325 0.5753336 0.765554759 3158.336
B_impute[10] 0.67563438 0.06162871 0.5714415 0.768452190 2902.297
B_impute[11] 0.67530152 0.06010048 0.5779499 0.768968719 2813.118
B_impute[12] 0.67451433 0.05874663 0.5801504 0.764538005 2611.613
Rhat4
a 1.0015286
bM 0.9990618
bB 1.0027464
p 0.9999588
theta 0.9993599
sigma 0.9997023
B_impute[1] 0.9989987
B_impute[2] 0.9991325
B_impute[3] 0.9991644
B_impute[4] 0.9992428
B_impute[5] 0.9994196
B_impute[6] 0.9990582
B_impute[7] 0.9990442
B_impute[8] 0.9986711
B_impute[9] 0.9994715
B_impute[10] 1.0000818
B_impute[11] 1.0001818
B_impute[12] 0.9988374
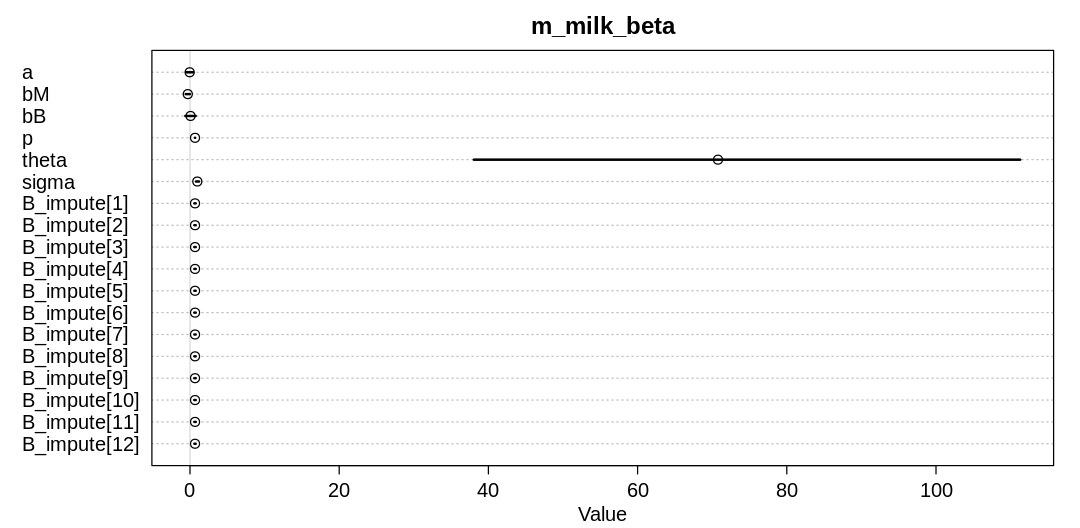
The theta parameter to the beta distribution is highly uncertain because there are simply too many
values that are compatible with these observations. The imputed values are essentially nothing more
than the mean of the non-NA observations of brain size. That is, any gentle tilt of the imputed
values we’d expect from using the observed mass seems to have disappeared.
Let’s zoom in on some parameters of interest:
iplot(function() {
plot(precis(m_milk_beta, depth=3), main="m_milk_beta", pars=c("a", "bM", "bB", "sigma"))
}, ar=4.5)
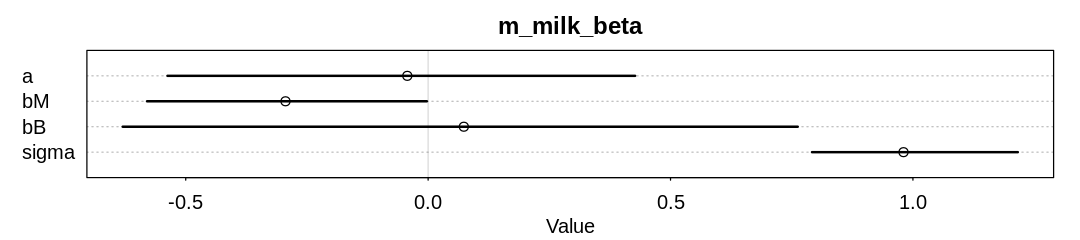
Compare these results to the figure produced from R code 15.20 in the chapter. The inference for
bM has shrunk even further. It’s likely bB has shrunk as well, though in this new model bB is
multiplied by a probability rather than a standardized value so we can expect it to be smaller.
15M3. Repeat the divorce data measurement error models, but this time double the standard errors. Can you explain how doubling the standard errors impacts inference?
Answer. Let’s start by reproducing the results from the chapter, for the sake of comparison:
data(WaffleDivorce)
d <- WaffleDivorce
dlist <- list(
D_obs = standardize(d$Divorce),
D_sd = d$Divorce.SE / sd(d$Divorce),
M = standardize(d$Marriage),
A = standardize(d$MedianAgeMarriage),
N = nrow(d)
)
m15.1 <- ulam(
alist(
D_obs ~ dnorm(D_true, D_sd),
vector[N]:D_true ~ dnorm(mu, sigma),
mu <- a + bA * A + bM * M,
a ~ dnorm(0, 0.2),
bA ~ dnorm(0, 0.5),
bM ~ dnorm(0, 0.5),
sigma ~ dexp(1)
),
cmdstan = TRUE, data = dlist, chains = 4, cores = 4
)
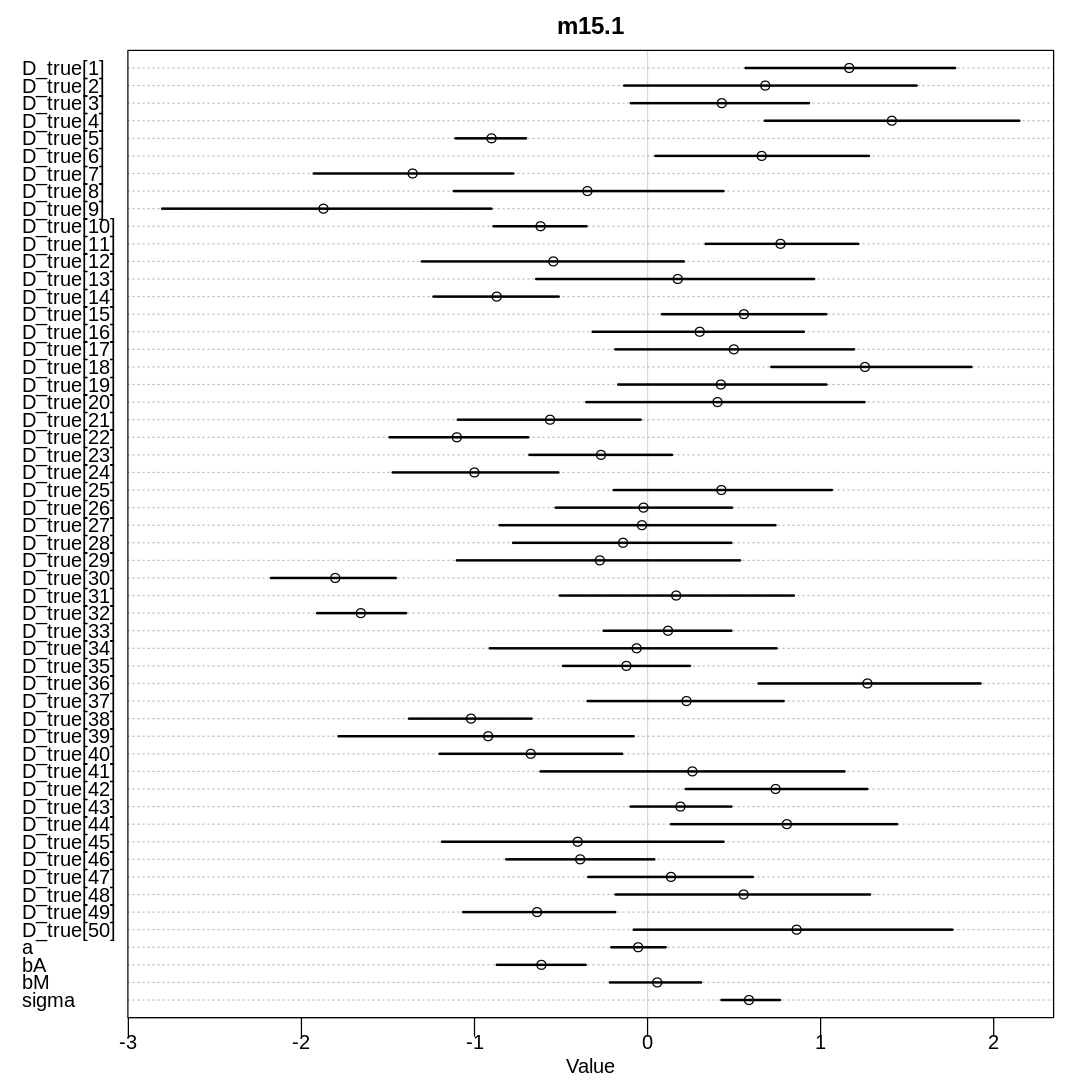
display(precis(m15.1, depth=3), mimetypes="text/plain")
iplot(function() {
plot(precis(m15.1, depth=3), main="m15.1")
}, ar=1)
dlist <- list(
D_obs = standardize(d$Divorce),
D_sd = d$Divorce.SE / sd(d$Divorce),
M_obs = standardize(d$Marriage),
M_sd = d$Marriage.SE / sd(d$Marriage),
A = standardize(d$MedianAgeMarriage),
N = nrow(d)
)
m15.2 <- ulam(
alist(
D_obs ~ dnorm(D_true, D_sd),
vector[N]:D_true ~ dnorm(mu, sigma),
mu <- a + bA * A + bM * M_true[i],
M_obs ~ dnorm(M_true, M_sd),
vector[N]:M_true ~ dnorm(0, 1),
a ~ dnorm(0, 0.2),
bA ~ dnorm(0, 0.5),
bM ~ dnorm(0, 0.5),
sigma ~ dexp(1)
),
cmdstan = TRUE, data = dlist, chains = 4, cores = 4
)
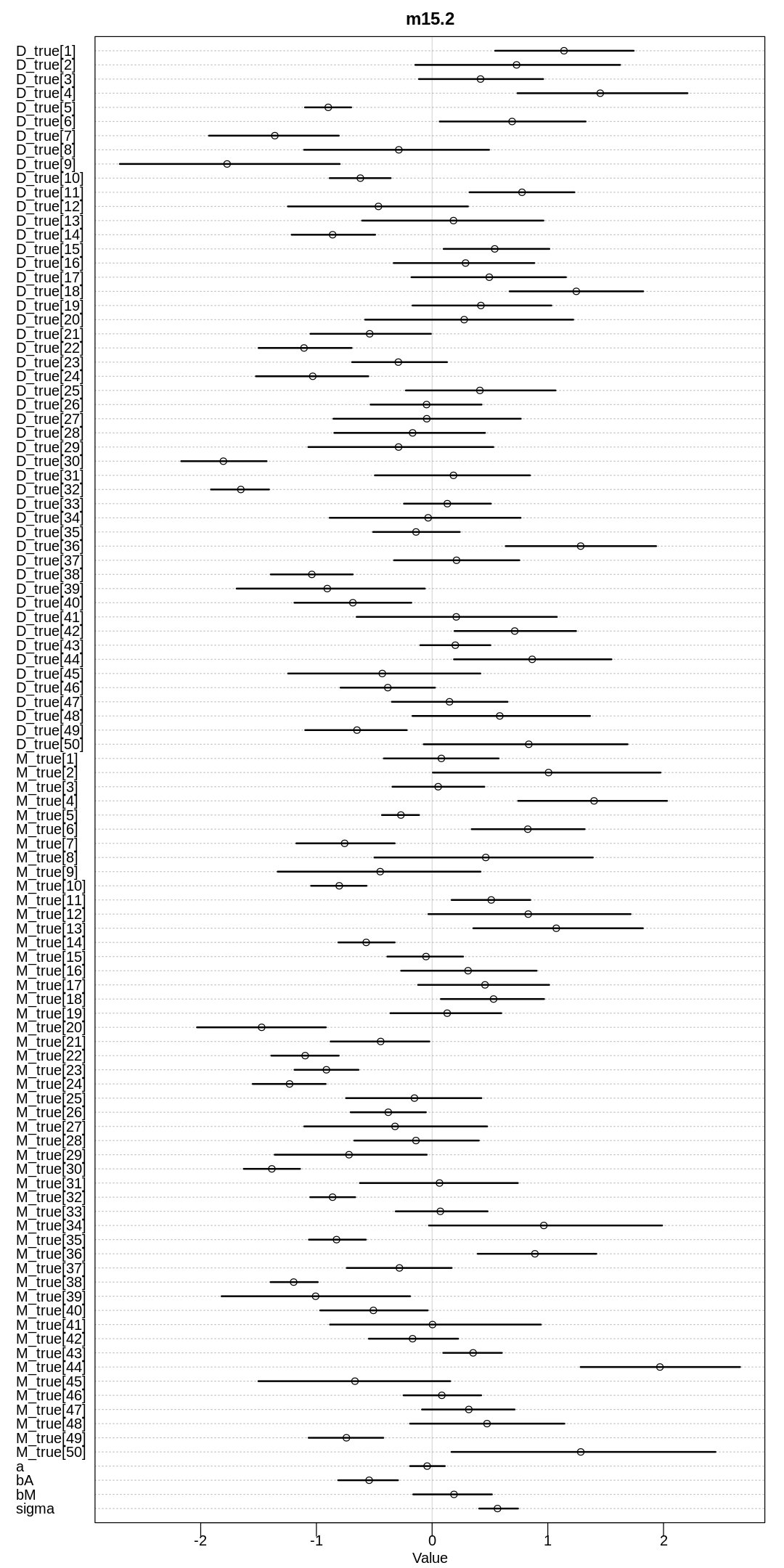
display(precis(m15.2, depth=3), mimetypes="text/plain")
iplot(function() {
plot(precis(m15.2, depth=3), main="m15.2")
}, ar=0.5)
Running MCMC with 4 parallel chains, with 1 thread(s) per chain...
Chain 1 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 1 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 1 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 1 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 1 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 1 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 1 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 1 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 2 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 2 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 2 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 2 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 2 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 2 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 2 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 3 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 3 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 3 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 3 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 3 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 3 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 3 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 3 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 3 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 3 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 3 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 4 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 4 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 4 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 4 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 4 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 4 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 4 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 4 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 1 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 1 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 1 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 1 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 1 finished in 0.2 seconds.
Chain 2 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 2 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 2 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 2 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 2 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 2 finished in 0.2 seconds.
Chain 3 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 3 finished in 0.2 seconds.
Chain 4 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 4 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 4 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 4 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 4 finished in 0.3 seconds.
All 4 chains finished successfully.
Mean chain execution time: 0.2 seconds.
Total execution time: 0.5 seconds.
mean sd 5.5% 94.5% n_eff Rhat4
D_true[1] 1.16529913 0.37670085 0.56646235 1.77686420 1826.9479 0.9997720
D_true[2] 0.68035140 0.53713821 -0.13498373 1.55454040 1763.5176 1.0022227
D_true[3] 0.42872481 0.33167476 -0.09605938 0.93261400 2683.5404 0.9995092
D_true[4] 1.41154159 0.46577940 0.67784049 2.14802785 2554.3427 0.9985109
D_true[5] -0.90160861 0.12550591 -1.10984500 -0.70248119 2592.0895 0.9990946
D_true[6] 0.65953298 0.39147481 0.04538031 1.27896865 2126.2524 1.0005978
D_true[7] -1.35749882 0.35406903 -1.92839990 -0.77569350 1818.2683 0.9992590
D_true[8] -0.34753309 0.48870640 -1.11989465 0.43857377 1775.4601 1.0015430
D_true[9] -1.87285832 0.60396528 -2.80459145 -0.90207235 1459.0806 1.0019322
D_true[10] -0.61835239 0.16858058 -0.89032025 -0.35274012 2657.2032 1.0005894
D_true[11] 0.76812978 0.27716226 0.33565835 1.21754440 2329.5430 0.9991595
D_true[12] -0.54472513 0.47509818 -1.30452415 0.20983636 1811.4371 1.0012212
D_true[13] 0.17434182 0.50165467 -0.64344098 0.96242766 1080.0249 1.0033488
D_true[14] -0.87181479 0.22336972 -1.23705925 -0.51341138 2245.2291 0.9999715
D_true[15] 0.55659024 0.30202896 0.08249217 1.03231220 2200.4551 0.9999940
D_true[16] 0.30132522 0.38326632 -0.31731629 0.90287237 2868.8095 0.9994837
D_true[17] 0.49857479 0.43252705 -0.18628874 1.19296880 3231.2556 0.9989415
D_true[18] 1.25611387 0.35837104 0.71500242 1.87129685 2111.6199 1.0001413
D_true[19] 0.42369517 0.38116782 -0.16956035 1.03324940 2269.2815 0.9993208
D_true[20] 0.40433882 0.51297364 -0.35406454 1.25353315 1176.9675 1.0014683
D_true[21] -0.56355219 0.33233896 -1.09647050 -0.04003655 2360.7146 0.9996213
D_true[22] -1.10207486 0.25450122 -1.49067225 -0.68943755 2347.1546 1.0005273
D_true[23] -0.26902316 0.26278468 -0.68275595 0.14115404 2774.6269 1.0001873
D_true[24] -1.00045957 0.29883294 -1.47296970 -0.51553198 1730.4090 1.0007305
D_true[25] 0.42674263 0.40008865 -0.19583232 1.06478540 2663.3554 0.9993704
D_true[26] -0.02343933 0.32142629 -0.53034520 0.48795575 2133.5721 1.0029464
D_true[27] -0.03251814 0.49720989 -0.85495812 0.73914262 2200.8593 0.9999593
D_true[28] -0.14182911 0.39517413 -0.77690007 0.48404311 2124.0202 1.0000422
D_true[29] -0.27579654 0.50811564 -1.10165660 0.53418449 2467.4737 0.9991335
D_true[30] -1.80465810 0.23055456 -2.17609660 -1.45517780 2087.2871 1.0008965
D_true[31] 0.16544667 0.43112034 -0.50715942 0.84543276 2966.5765 0.9993029
D_true[32] -1.65694118 0.16025453 -1.90951420 -1.39546590 2298.8496 1.0002954
D_true[33] 0.11802736 0.23494846 -0.25410918 0.48409419 2850.4477 1.0001158
D_true[34] -0.06339266 0.51499171 -0.91162970 0.74493650 1419.7932 1.0014848
D_true[35] -0.12244562 0.22985512 -0.48932804 0.24497560 2508.5167 0.9996304
D_true[36] 1.27047247 0.40131977 0.64126050 1.92421385 2318.8039 0.9998624
D_true[37] 0.22497356 0.35980983 -0.34656786 0.78629118 2595.7440 0.9991507
D_true[38] -1.02029160 0.22501755 -1.37862575 -0.67071761 2704.1224 0.9993599
D_true[39] -0.92104471 0.53912631 -1.78547790 -0.08014792 1824.4961 0.9995028
D_true[40] -0.67496281 0.33746049 -1.20182360 -0.14724809 2433.9734 1.0004294
D_true[41] 0.25872653 0.54692102 -0.61817572 1.13789520 2040.4956 0.9988532
D_true[42] 0.73961065 0.33014936 0.22117780 1.26946220 2414.9662 0.9990285
D_true[43] 0.18995112 0.18223568 -0.09803334 0.48426666 2414.6235 1.0000140
D_true[44] 0.80465016 0.41707592 0.13549761 1.44203035 1351.6125 1.0031868
D_true[45] -0.40386974 0.51067738 -1.18794350 0.43917189 1909.1625 0.9988625
D_true[46] -0.38938278 0.26783573 -0.81655354 0.03900730 2385.6420 0.9999969
D_true[47] 0.13534855 0.29424111 -0.34327577 0.60873387 2989.7362 0.9993935
D_true[48] 0.55514460 0.46916305 -0.18476896 1.28536950 3325.5506 0.9983973
D_true[49] -0.63843072 0.27863705 -1.06624065 -0.18693361 2526.1883 1.0011626
D_true[50] 0.86124113 0.59606065 -0.07979191 1.76178105 1617.5362 1.0017955
a -0.05446510 0.09751551 -0.20976111 0.10457820 1386.6334 1.0004440
bA -0.61351468 0.16049184 -0.87111548 -0.35784110 900.5241 1.0025891
bM 0.05598574 0.16706251 -0.21729289 0.30902512 873.6370 1.0018299
sigma 0.58590288 0.10778943 0.42688904 0.76552462 486.0034 1.0095561
Running MCMC with 4 parallel chains, with 1 thread(s) per chain...
Chain 1 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 1 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 1 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 1 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 2 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 2 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 2 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 2 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 2 Exception: normal_lpdf: Scale parameter is 0, but must be positive! (in '/tmp/RtmpjPjDJw/model-122086d758.stan', line 28, column 4 to column 34)
Chain 2 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 2 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 2
Chain 3 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 3 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 3 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 3 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 3 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 3 Exception: normal_lpdf: Scale parameter is 0, but must be positive! (in '/tmp/RtmpjPjDJw/model-122086d758.stan', line 28, column 4 to column 34)
Chain 3 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 3 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 3
Chain 4 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 4 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 4 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 4 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 1 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 1 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 1 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 1 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 1 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 1 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 1 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 2 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 2 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 2 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 2 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 2 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 3 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 3 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 3 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 3 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 3 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 4 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 4 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 4 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 4 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 1 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 2 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 2 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 2 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 3 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 3 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 3 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 4 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 4 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 4 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 1 finished in 0.4 seconds.
Chain 3 finished in 0.3 seconds.
Chain 2 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 4 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 2 finished in 0.4 seconds.
Chain 4 finished in 0.4 seconds.
All 4 chains finished successfully.
Mean chain execution time: 0.4 seconds.
Total execution time: 0.6 seconds.
mean sd 5.5% 94.5% n_eff Rhat4
D_true[1] 1.13980066 0.3672073 0.54473938 1.7421660 2020.261 0.9991496
D_true[2] 0.72989332 0.5503889 -0.14497435 1.6243327 2388.551 0.9985643
D_true[3] 0.41832269 0.3381948 -0.11368834 0.9595620 2588.476 0.9999522
D_true[4] 1.45304387 0.4546278 0.73780377 2.2073164 1917.647 1.0003916
D_true[5] -0.89750179 0.1241661 -1.09859765 -0.6993325 2744.301 1.0005105
D_true[6] 0.69134812 0.3979001 0.06562878 1.3267263 2694.820 0.9993062
D_true[7] -1.35943323 0.3459120 -1.92849000 -0.8064240 2797.510 0.9995818
D_true[8] -0.28779986 0.4916344 -1.10661110 0.4921008 2049.506 1.0009338
D_true[9] -1.77196219 0.6044402 -2.69894100 -0.7984069 1621.656 1.0006758
D_true[10] -0.62027647 0.1645052 -0.88511736 -0.3583054 3348.679 0.9988624
D_true[11] 0.77699272 0.2810548 0.32338724 1.2301131 2022.770 0.9989415
D_true[12] -0.46395623 0.4917993 -1.24678390 0.3100883 1947.360 0.9988687
D_true[13] 0.18466706 0.4984056 -0.60521819 0.9620114 1417.958 0.9987119
D_true[14] -0.85982422 0.2266755 -1.21400160 -0.4921531 2480.290 1.0002637
D_true[15] 0.54116474 0.2880502 0.09933228 1.0134607 3103.837 0.9999754
D_true[16] 0.28906162 0.3791991 -0.33203928 0.8832713 2187.039 0.9993098
D_true[17] 0.49396919 0.4189143 -0.17879638 1.1568778 2478.119 0.9986335
D_true[18] 1.24608606 0.3573026 0.66986521 1.8242623 2686.657 1.0001980
D_true[19] 0.42154247 0.3783306 -0.16917162 1.0305995 2987.535 1.0002947
D_true[20] 0.27708897 0.5574021 -0.57884490 1.2197770 1252.782 0.9987722
D_true[21] -0.53963402 0.3207615 -1.05170550 -0.0102739 3177.273 0.9993598
D_true[22] -1.10708126 0.2516425 -1.50025465 -0.6952419 2456.320 0.9991992
D_true[23] -0.29196223 0.2548001 -0.69192237 0.1285582 2615.308 0.9985269
D_true[24] -1.03161297 0.3008844 -1.52290475 -0.5511805 2632.526 0.9985800
D_true[25] 0.41285163 0.4124108 -0.22655639 1.0666809 3259.927 0.9983697
D_true[26] -0.04792652 0.3034968 -0.53220768 0.4272377 2364.137 0.9996221
D_true[27] -0.04623642 0.5140159 -0.85306116 0.7662948 2589.839 0.9988235
D_true[28] -0.16844509 0.3949712 -0.84551703 0.4571312 1849.513 1.0019656
D_true[29] -0.28993034 0.5012341 -1.06958630 0.5291909 2292.606 0.9985251
D_true[30] -1.80438402 0.2319326 -2.16829345 -1.4303780 1944.205 1.0026937
⋮ ⋮ ⋮ ⋮ ⋮ ⋮ ⋮
M_true[25] -0.152868519 0.36721297 -0.74386089 0.42564432 3721.3290 0.9990446
M_true[26] -0.378903023 0.20604708 -0.70358575 -0.05383121 3270.8281 0.9989264
M_true[27] -0.320480175 0.49159103 -1.10519135 0.47542391 2717.3530 1.0002243
M_true[28] -0.139549026 0.34161527 -0.67233812 0.40379155 2575.1460 0.9993844
M_true[29] -0.718199643 0.42032015 -1.36067610 -0.04640796 3469.1455 0.9989717
M_true[30] -1.385296362 0.15555509 -1.62810960 -1.14155055 2945.3393 1.0006579
M_true[31] 0.064082656 0.43066441 -0.62428938 0.74098219 2936.6891 0.9992723
M_true[32] -0.860433145 0.12348629 -1.05343770 -0.66506229 2970.6106 0.9998388
M_true[33] 0.071180967 0.24761218 -0.31449523 0.47968621 2977.6313 0.9992579
M_true[34] 0.965363880 0.62923413 -0.02728858 1.98761395 2560.6416 0.9983876
M_true[35] -0.826105916 0.15538014 -1.06481480 -0.57270177 3231.9187 0.9998117
M_true[36] 0.888540910 0.32103142 0.39403025 1.41934490 2522.7284 0.9992918
M_true[37] -0.282542098 0.28316057 -0.73776220 0.16931516 2917.1523 1.0009205
M_true[38] -1.196606841 0.12610830 -1.39632930 -0.98794782 2274.0078 0.9994018
M_true[39] -1.006728396 0.50639594 -1.81955690 -0.19055231 2300.9962 1.0021731
M_true[40] -0.507620174 0.29019788 -0.96763578 -0.03805253 2848.2815 1.0002209
M_true[41] 0.003610646 0.57901820 -0.88233998 0.93984612 2575.4191 0.9998823
M_true[42] -0.168825699 0.23540747 -0.54638417 0.22476800 3068.1449 0.9990647
M_true[43] 0.354329161 0.16036737 0.09608789 0.60301057 3339.1457 0.9991328
M_true[44] 1.968909632 0.42897395 1.28348095 2.66120435 2090.2837 0.9992804
M_true[45] -0.666842085 0.52079232 -1.50119905 0.15627512 2788.2095 0.9989813
M_true[46] 0.083925491 0.21217465 -0.24694421 0.42397530 2487.9784 0.9991141
M_true[47] 0.317316058 0.25239524 -0.08739298 0.71246712 2623.8162 0.9998099
M_true[48] 0.474560523 0.41411576 -0.19063167 1.14342125 2243.7097 0.9988102
M_true[49] -0.740652101 0.20116619 -1.06708950 -0.42189996 2876.6849 0.9992264
M_true[50] 1.284739162 0.71606319 0.16746741 2.44926415 2683.6225 0.9987561
a -0.042854779 0.09496423 -0.19077060 0.11008004 1648.8423 0.9996295
bA -0.544299049 0.16209833 -0.81209181 -0.29511154 996.5373 1.0028343
bM 0.189021324 0.20999733 -0.16315066 0.51688197 695.7240 1.0021214
sigma 0.565245555 0.10878059 0.40696218 0.74226751 562.8355 1.0015088
Doubling the standard error:
d$doubleSE = 2*d$Divorce.SE
dlist <- list(
D_obs = standardize(d$Divorce),
D_sd = d$doubleSE / sd(d$Divorce),
M = standardize(d$Marriage),
A = standardize(d$MedianAgeMarriage),
N = nrow(d)
)
m.double.se.1 <- ulam(
alist(
D_obs ~ dnorm(D_true, D_sd),
vector[N]:D_true ~ dnorm(mu, sigma),
mu <- a + bA * A + bM * M,
a ~ dnorm(0, 0.2),
bA ~ dnorm(0, 0.5),
bM ~ dnorm(0, 0.5),
sigma ~ dexp(1)
),
cmdstan = TRUE, data = dlist, chains = 4, cores = 4
)
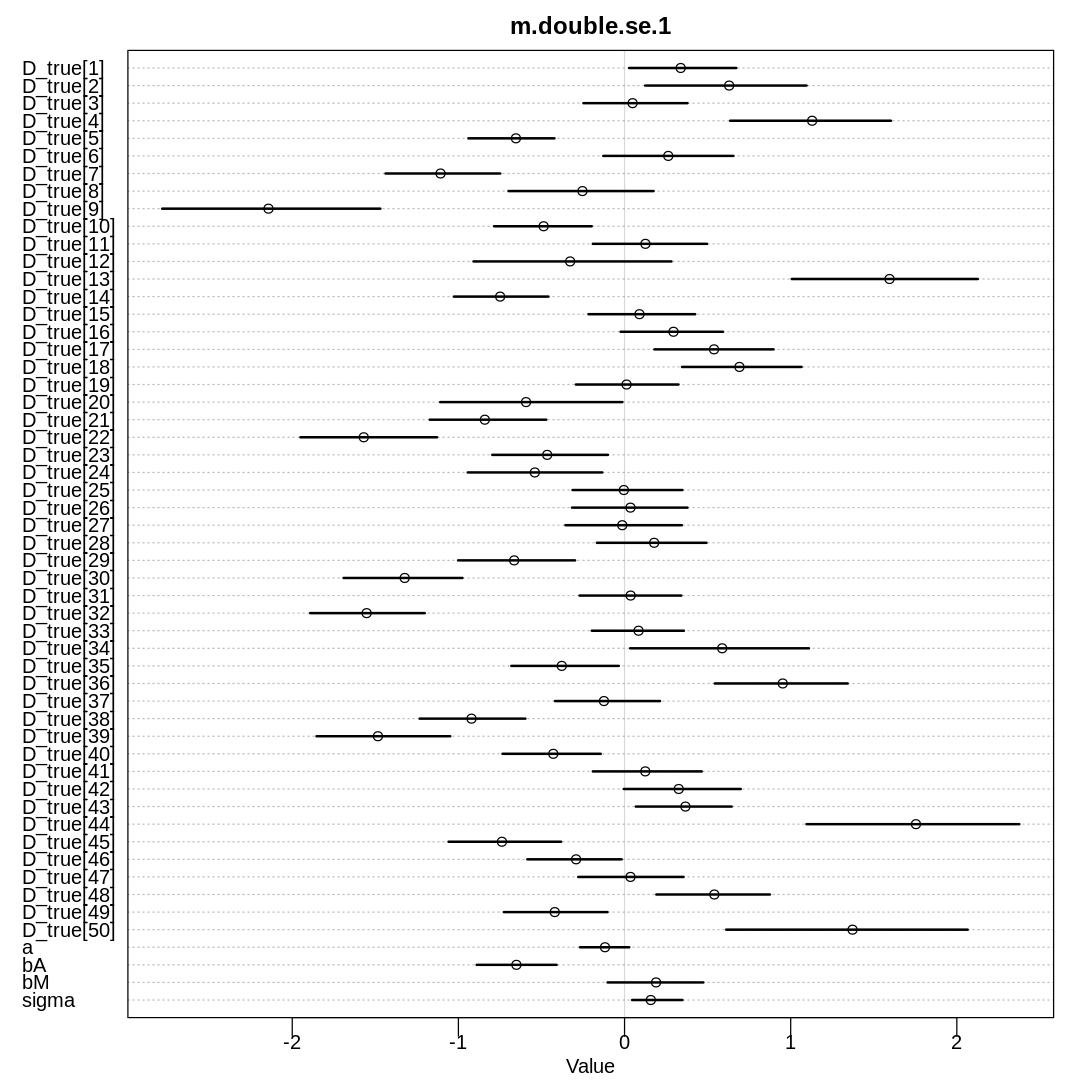
display(precis(m.double.se.1, depth=3), mimetypes="text/plain")
iplot(function() {
plot(precis(m.double.se.1, depth=3), main="m.double.se.1")
}, ar=1)
dlist <- list(
D_obs = standardize(d$Divorce),
D_sd = d$doubleSE / sd(d$Divorce),
M_obs = standardize(d$Marriage),
M_sd = d$Marriage.SE / sd(d$Marriage),
A = standardize(d$MedianAgeMarriage),
N = nrow(d)
)
m.double.se.2 <- ulam(
alist(
D_obs ~ dnorm(D_true, D_sd),
vector[N]:D_true ~ dnorm(mu, sigma),
mu <- a + bA * A + bM * M_true[i],
M_obs ~ dnorm(M_true, M_sd),
vector[N]:M_true ~ dnorm(0, 1),
a ~ dnorm(0, 0.2),
bA ~ dnorm(0, 0.5),
bM ~ dnorm(0, 0.5),
sigma ~ dexp(1)
),
cmdstan = TRUE, data = dlist, chains = 4, cores = 4
)
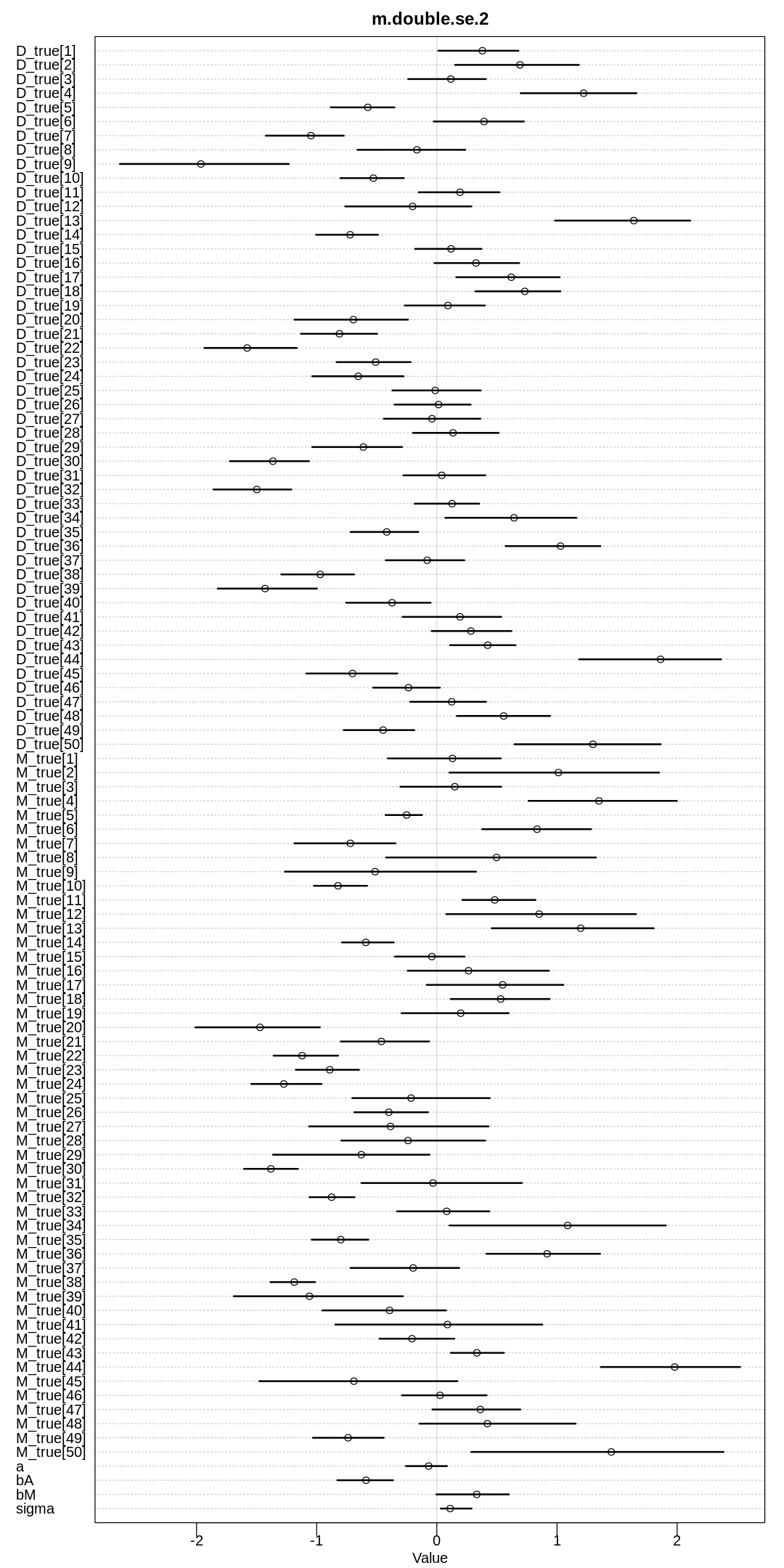
display(precis(m.double.se.2, depth=3), mimetypes="text/plain")
iplot(function() {
plot(precis(m.double.se.2, depth=3), main="m.double.se.2")
}, ar=0.5)
Running MCMC with 4 parallel chains, with 1 thread(s) per chain...
Chain 1 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 1 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 1 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 2 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 2 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 2 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 3 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 3 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 3 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 4 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 4 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 1 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 1 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 2 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 2 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 3 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 3 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 4 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 4 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 1 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 1 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 2 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 2 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 3 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 3 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 3 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 3 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 3 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 4 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 1 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 3 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 3 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 4 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 4 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 4 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 3 finished in 0.4 seconds.
Chain 1 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 4 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 4 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 1 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 4 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 1 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 2 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 4 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 4 finished in 0.7 seconds.
Chain 1 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 1 finished in 0.8 seconds.
Chain 2 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 2 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 2 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 2 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 2 finished in 2.2 seconds.
All 4 chains finished successfully.
Mean chain execution time: 1.0 seconds.
Total execution time: 2.2 seconds.
Warning: 26 of 2000 (1.0%) transitions ended with a divergence.
This may indicate insufficient exploration of the posterior distribution.
Possible remedies include:
* Increasing adapt_delta closer to 1 (default is 0.8)
* Reparameterizing the model (e.g. using a non-centered parameterization)
* Using informative or weakly informative prior distributions
22 of 2000 (1.0%) transitions hit the maximum treedepth limit of 10 or 2^10-1 leapfrog steps.
Trajectories that are prematurely terminated due to this limit will result in slow exploration.
Increasing the max_treedepth limit can avoid this at the expense of more computation.
If increasing max_treedepth does not remove warnings, try to reparameterize the model.
mean sd 5.5% 94.5% n_eff Rhat4
D_true[1] 0.33796572 0.21582894 0.027589873 0.67252494 236.76301 1.014180
D_true[2] 0.62947793 0.31168698 0.124353785 1.09543480 220.40308 1.020648
D_true[3] 0.04841299 0.20678361 -0.247430415 0.37882753 549.05169 1.002715
D_true[4] 1.12905704 0.31365612 0.635662415 1.60328915 214.67947 1.026316
D_true[5] -0.65377791 0.16034456 -0.939621315 -0.42165778 173.23045 1.023178
D_true[6] 0.26316311 0.24978849 -0.128048560 0.65538663 306.61900 1.009790
D_true[7] -1.10742594 0.21732411 -1.438743350 -0.74813529 430.64113 1.011535
D_true[8] -0.25339828 0.28199675 -0.698303350 0.17495354 255.05753 1.013457
D_true[9] -2.14312320 0.41295933 -2.782816350 -1.46995160 206.28805 1.019418
D_true[10] -0.48705608 0.18567907 -0.785085160 -0.19763261 465.86484 1.008968
D_true[11] 0.12557759 0.22536052 -0.190852545 0.49629107 233.26458 1.014086
D_true[12] -0.32751822 0.37311658 -0.908953875 0.28187891 229.07483 1.014311
D_true[13] 1.59482665 0.35063885 1.006406650 2.12598575 138.94207 1.027431
D_true[14] -0.74863071 0.18364053 -1.027192450 -0.45797249 725.91562 1.004536
D_true[15] 0.08982116 0.20895956 -0.217009440 0.42511147 323.31161 1.004783
D_true[16] 0.29448084 0.20254489 -0.023449970 0.59243579 359.42212 1.003181
D_true[17] 0.53855027 0.23803871 0.179682630 0.89699193 372.93201 1.005101
D_true[18] 0.69156478 0.23760357 0.345972585 1.06529955 219.43478 1.015470
D_true[19] 0.01196021 0.20513287 -0.292825305 0.32458850 481.09538 1.001699
D_true[20] -0.59295331 0.35022430 -1.109419600 -0.01348071 259.99886 1.012581
D_true[21] -0.84093494 0.22636292 -1.172218850 -0.47132749 410.37739 1.005227
D_true[22] -1.56966665 0.26253209 -1.950601650 -1.12831050 182.68226 1.022074
D_true[23] -0.46547881 0.22051994 -0.796227160 -0.10014746 368.72365 1.007733
D_true[24] -0.53996588 0.25540117 -0.943720300 -0.13405369 262.57847 1.020685
D_true[25] -0.00404805 0.20885334 -0.313141805 0.34816536 462.60509 1.001810
D_true[26] 0.03556651 0.21453105 -0.316112430 0.37854043 414.39580 1.003293
D_true[27] -0.01375848 0.22722340 -0.355973710 0.34481386 434.48602 1.001883
D_true[28] 0.17821194 0.21297761 -0.166225005 0.49309792 323.32638 1.006560
D_true[29] -0.66441592 0.23032608 -1.001813750 -0.29779869 483.74924 1.007189
D_true[30] -1.32330272 0.23572433 -1.690969800 -0.97576939 202.44262 1.034145
D_true[31] 0.03665304 0.20061424 -0.270965560 0.34164883 481.31775 1.002319
D_true[32] -1.55229859 0.21759792 -1.892036100 -1.20229905 228.69233 1.028532
D_true[33] 0.08431884 0.17853591 -0.197324755 0.35734198 468.68184 1.002113
D_true[34] 0.58767295 0.34871339 0.033809024 1.10981235 228.57250 1.019512
D_true[35] -0.37795638 0.20634101 -0.681413805 -0.03474520 312.20969 1.006037
D_true[36] 0.95217628 0.25611169 0.542974750 1.34295930 200.12218 1.018736
D_true[37] -0.12412461 0.20532204 -0.418619655 0.21359914 480.65174 1.000627
D_true[38] -0.92107028 0.19898630 -1.233116650 -0.59688532 313.20667 1.019827
D_true[39] -1.48428354 0.26331386 -1.853833450 -1.04920445 286.03251 1.019377
D_true[40] -0.42893058 0.19022471 -0.734642685 -0.14324799 706.00854 1.004737
D_true[41] 0.12493998 0.21404886 -0.190648675 0.46486038 438.52776 1.004675
D_true[42] 0.32613166 0.22839459 -0.004690787 0.69938573 275.37900 1.006487
D_true[43] 0.36641722 0.18829718 0.068024864 0.64484670 193.80211 1.015689
D_true[44] 1.75382435 0.40725423 1.095382350 2.37603800 146.49939 1.032014
D_true[45] -0.73812669 0.23060977 -1.058904000 -0.38183290 409.10015 1.009005
D_true[46] -0.29157763 0.19121975 -0.585240370 -0.01712644 542.94456 1.001173
D_true[47] 0.03571322 0.20550448 -0.278653425 0.35532445 411.69635 1.003303
D_true[48] 0.54032926 0.22679137 0.191324580 0.87468402 285.35007 1.009658
D_true[49] -0.42001603 0.20432710 -0.726107710 -0.10278328 490.55769 1.008375
D_true[50] 1.37216976 0.46116025 0.611850845 2.06541330 173.19466 1.030323
a -0.11762847 0.09254747 -0.268001550 0.02835270 154.73145 1.002047
bA -0.65083652 0.15185027 -0.889979940 -0.40885810 163.46987 1.019597
bM 0.18958345 0.18231647 -0.102041000 0.47425205 187.29157 1.021093
sigma 0.15797996 0.09803259 0.045619303 0.34843333 40.21924 1.110314
Running MCMC with 4 parallel chains, with 1 thread(s) per chain...
Chain 1 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 1 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 1 Exception: normal_lpdf: Scale parameter is 0, but must be positive! (in '/tmp/RtmpjPjDJw/model-12fa46c90.stan', line 28, column 4 to column 34)
Chain 1 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 1 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 1
Chain 2 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 3 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 3 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 4 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 4 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 4 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 4 Exception: normal_lpdf: Scale parameter is 0, but must be positive! (in '/tmp/RtmpjPjDJw/model-12fa46c90.stan', line 28, column 4 to column 34)
Chain 4 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 4 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 4
Chain 2 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 4 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 2 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 3 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 4 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 1 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 3 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 2 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 3 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 4 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 1 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 4 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 4 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 3 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 3 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 4 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 4 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 4 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 4 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 4 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 4 finished in 0.7 seconds.
Chain 1 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 1 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 3 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 2 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 3 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 3 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 1 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 1 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 3 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 1 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 2 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 2 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 3 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 3 finished in 1.3 seconds.
Chain 1 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 1 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 2 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 1 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 1 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 2 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 1 finished in 1.9 seconds.
Chain 2 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 2 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 2 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 2 finished in 2.8 seconds.
All 4 chains finished successfully.
Mean chain execution time: 1.7 seconds.
Total execution time: 2.9 seconds.
Warning: 445 of 2000 (22.0%) transitions ended with a divergence.
This may indicate insufficient exploration of the posterior distribution.
Possible remedies include:
* Increasing adapt_delta closer to 1 (default is 0.8)
* Reparameterizing the model (e.g. using a non-centered parameterization)
* Using informative or weakly informative prior distributions
mean sd 5.5% 94.5% n_eff Rhat4
D_true[1] 0.37953293 0.2398603 0.01237775 0.6787418 9.805153 1.146924
D_true[2] 0.69196627 0.3523082 0.15158532 1.1824500 12.469865 1.123998
D_true[3] 0.11663164 0.2254270 -0.23885513 0.4072780 6.653921 1.230899
D_true[4] 1.22158311 0.3119979 0.69807085 1.6619734 66.510479 1.067652
D_true[5] -0.57465848 0.1764928 -0.88366910 -0.3530077 7.000977 1.195604
D_true[6] 0.39357630 0.2683339 -0.02626940 0.7254170 13.364836 1.138957
D_true[7] -1.04783714 0.2253261 -1.42403550 -0.7743593 12.841747 1.116780
D_true[8] -0.16575146 0.3102746 -0.66099346 0.2364567 12.702190 1.116254
D_true[9] -1.96373960 0.4200831 -2.63845265 -1.2318186 400.513116 1.013574
D_true[10] -0.52815005 0.1677250 -0.80202447 -0.2736530 390.916477 1.010018
D_true[11] 0.19327236 0.2187164 -0.15183088 0.5218317 200.169074 1.036598
D_true[12] -0.20145125 0.3498259 -0.76296994 0.2873224 16.480917 1.109628
D_true[13] 1.63929540 0.3871895 0.98354415 2.1092200 7.252962 1.209580
D_true[14] -0.72190047 0.1711595 -1.00516870 -0.4894583 19.605170 1.087852
D_true[15] 0.11847032 0.1899755 -0.18137113 0.3714890 12.804508 1.141989
D_true[16] 0.32601672 0.2262484 -0.02023291 0.6848860 88.587983 1.037856
D_true[17] 0.61935379 0.2875835 0.16176984 1.0222969 5.665619 1.287454
D_true[18] 0.73238124 0.2407032 0.32078948 1.0285387 15.077739 1.106224
D_true[19] 0.09252788 0.2342369 -0.26708202 0.3998111 7.494055 1.211244
D_true[20] -0.69459447 0.2860700 -1.18549245 -0.2400368 264.378090 1.014034
D_true[21] -0.81075193 0.2038724 -1.13111530 -0.4978065 408.929931 1.016963
D_true[22] -1.57889329 0.2371380 -1.93478115 -1.1654505 421.561441 1.016519
D_true[23] -0.50900328 0.1986487 -0.83406531 -0.2181990 133.318558 1.035616
D_true[24] -0.65350689 0.2366713 -1.03768115 -0.2767566 309.108215 1.015143
D_true[25] -0.01350890 0.2223810 -0.37072281 0.3659846 483.597192 1.012460
D_true[26] 0.01433202 0.2046745 -0.35245535 0.2819965 166.364248 1.033950
D_true[27] -0.04005472 0.2447914 -0.44188560 0.3629682 570.376649 1.009484
D_true[28] 0.13487870 0.2242061 -0.19848964 0.5155723 353.172665 1.010019
D_true[29] -0.61159603 0.2547695 -1.03730385 -0.2895010 11.250510 1.138764
D_true[30] -1.36428192 0.2051555 -1.72247665 -1.0638615 591.624331 1.004585
⋮ ⋮ ⋮ ⋮ ⋮ ⋮ ⋮
M_true[25] -0.21450375 0.37080789 -0.704379665 0.44097338 67.650780 1.052598
M_true[26] -0.39950867 0.19107744 -0.686060750 -0.07300721 246.048323 1.018746
M_true[27] -0.38521611 0.48143315 -1.063819300 0.43026703 39.630029 1.060365
M_true[28] -0.23981025 0.39151857 -0.795504935 0.40261822 8.980552 1.149880
M_true[29] -0.62854469 0.44034875 -1.364395950 -0.06096430 11.298713 1.130044
M_true[30] -1.38141541 0.13872286 -1.605390000 -1.15659475 913.607736 1.011502
M_true[31] -0.03150485 0.43527563 -0.626935000 0.70815564 15.153782 1.093691
M_true[32] -0.87557438 0.11473507 -1.060731000 -0.68501714 833.805459 1.009311
M_true[33] 0.08227647 0.23253696 -0.332239370 0.43675595 641.999055 1.007397
M_true[34] 1.08883925 0.57306156 0.104921880 1.90639830 48.956544 1.054365
M_true[35] -0.79911565 0.14882447 -1.041555950 -0.56968936 287.250840 1.008154
M_true[36] 0.91791597 0.29856559 0.411716810 1.35920220 400.275892 1.014239
M_true[37] -0.19707880 0.30266146 -0.717625440 0.18640400 7.613847 1.173246
M_true[38] -1.18629699 0.11594019 -1.384891150 -1.01389470 187.089798 1.025461
M_true[39] -1.06061164 0.43551755 -1.690460850 -0.28232032 179.154085 1.020598
M_true[40] -0.39334273 0.34190572 -0.952861170 0.07671980 7.042950 1.199141
M_true[41] 0.08824213 0.54307125 -0.844324385 0.87879552 54.645211 1.040951
M_true[42] -0.20635177 0.21146470 -0.477607215 0.14519977 15.547123 1.089552
M_true[43] 0.33338974 0.13822960 0.115647790 0.55802736 2154.259473 1.000542
M_true[44] 1.97973930 0.36420964 1.364875800 2.52349315 623.179581 1.004525
M_true[45] -0.68998058 0.50541114 -1.477439800 0.17077350 1107.339486 1.004155
M_true[46] 0.02659864 0.23250230 -0.290543000 0.41404550 8.907110 1.196804
M_true[47] 0.36215763 0.23708300 -0.037589007 0.69513531 55.666390 1.047142
M_true[48] 0.42050286 0.40579333 -0.144471320 1.15544260 62.716890 1.041190
M_true[49] -0.73834201 0.18019941 -1.031957950 -0.44189984 900.957758 1.004596
M_true[50] 1.45234177 0.66575837 0.286931630 2.38665805 52.409417 1.054564
a -0.06795557 0.11847845 -0.257483800 0.08447670 5.230863 1.321037
bA -0.58923142 0.13619825 -0.828265435 -0.36454046 312.111245 1.020398
bM 0.33241402 0.18764196 -0.003598285 0.59993062 68.101948 1.071576
sigma 0.11124654 0.09173301 0.033021518 0.28845468 17.889911 1.152191
Comparing the most important parameters:
iplot(function() {
plot(
coeftab(m.double.se.2, m15.2, m.double.se.1, m15.1), pars=c("a", "bA", "bM", "sigma"),
main="All models"
)
}, ar=1.7)
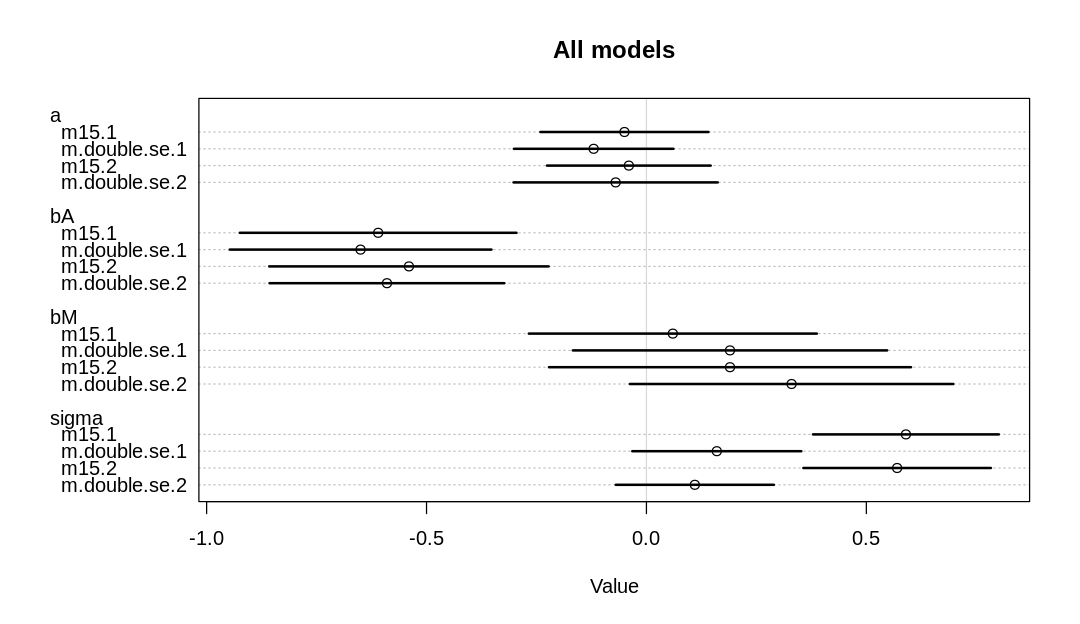
Notice we are getting some divergent transitions. It’s likely we’re running into these issues because there is more parameter space to explore; we’re less certain about all our inferences and so we need to cover more possibilities. Like vague priors, this can lead to divergent transitions.
The a, bA, and bM parameters have increased. With less certainty in the measurements, it’s
more plausible that a stronger relationship holds (similar to what we saw in Figure 15.3). In
contrast, the sigma parameter has decreased, because we can now explain more variation in terms of
measurement error than general noise.
We also see, in general, more uncertainty in parameter estimates across the board (wider HDPI bars).
15M4. Simulate data from this DAG: \(X \rightarrow Y \rightarrow Z\). Now fit a model that predicts \(Y\) using both \(X\) and \(Z\). What kind of confound arises, in terms of inferring the causal influence of \(X\) on \(Y\)?
Answer. First lets simulate, with normal distributions:
N <- 1000
X <- rnorm(N)
Y <- rnorm(N, mean=X)
Z <- rnorm(N, mean=Y)
dat_list <- list(
X = X,
Y = Y,
Z = Z
)
m_confound <- ulam(
alist(
Y ~ dnorm(mu, sigma),
mu <- a + bX*X + bZ*Z,
c(a, bX, bZ) ~ dnorm(0, 1),
sigma ~ dexp(1)
), cmdstan = TRUE, data=dat_list, chains=4, cores=4
)
display(precis(m_confound, depth=3), mimetypes="text/plain")
iplot(function() {
plot(precis(m_confound, depth=3), main="m_confound")
}, ar=4.5)
Running MCMC with 4 parallel chains, with 1 thread(s) per chain...
Chain 1 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 1 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 1 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 1 Exception: normal_lpdf: Scale parameter is 0, but must be positive! (in '/tmp/RtmpjPjDJw/model-122e9e7fe8.stan', line 21, column 4 to column 29)
Chain 1 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 1 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 1
Chain 2 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 2 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 3 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 3 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 4 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 4 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 1 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 1 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 2 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 2 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 3 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 3 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 4 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 4 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 1 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 2 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 3 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 4 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 1 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 1 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 1 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 2 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 2 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 3 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 3 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 4 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 4 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 1 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 2 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 2 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 3 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 4 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 1 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 1 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 2 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 3 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 4 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 4 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 1 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 2 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 3 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 4 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 1 finished in 0.8 seconds.
Chain 2 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 3 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 3 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 4 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 2 finished in 0.8 seconds.
Chain 3 finished in 0.9 seconds.
Chain 4 finished in 0.8 seconds.
All 4 chains finished successfully.
Mean chain execution time: 0.8 seconds.
Total execution time: 0.9 seconds.
mean sd 5.5% 94.5% n_eff Rhat4
bZ 0.4915055 0.01581945 0.46620708 0.51615314 1370.006 1.001106
bX 0.5081546 0.02644927 0.46587723 0.55098527 1230.789 1.002709
a -0.0011672 0.02145382 -0.03532943 0.03244627 1503.938 1.001742
sigma 0.6913835 0.01513483 0.66766894 0.71596408 1687.319 1.001510
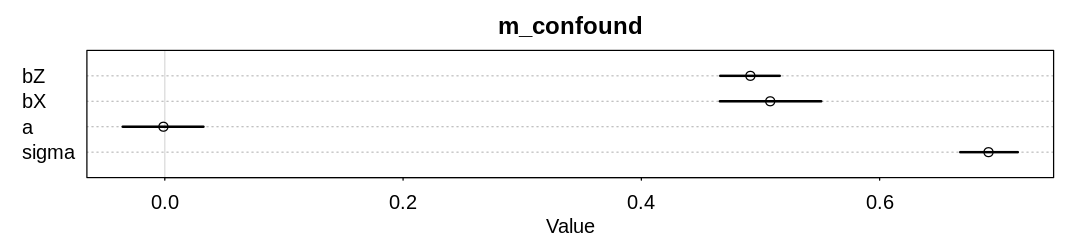

The confound we’re running into is post-treatment bias (conditioning on the outcome).
In this case \(Z\) is a better predictor of \(Y\) than \(X\) (notice the credible intervals). This is because the function between \(X\) and \(Y\) is the same as the function between \(Y\) and \(Z\), specifically the normal distribution with standard deviation equal one. Since \(Y\) will take on more dispersed values than \(X\), and \(Z\) more dispersed values than \(Y\), the more widely dispersed values of \(Z\) will predict \(Y\) better.
15M5. Return to the singing bird model, m15.9, and compare the posterior estimates of cat
presence (PrC1) to the true simulated values. How good is the model at inferring the missing data?
Can you think of a way to change the simulation so that the precision of the inference is stronger?
set.seed(9)
N_houses <- 100L
alpha <- 5
beta <- (-3)
k <- 0.5
r <- 0.2
cat <- rbern(N_houses, k)
notes <- rpois(N_houses, alpha + beta * cat)
R_C <- rbern(N_houses, r)
cat_obs <- cat
cat_obs[R_C == 1] <- (-9L)
dat <- list(
notes = notes,
cat = cat_obs,
RC = R_C,
N = as.integer(N_houses)
)
m15.9 <- ulam(
alist(
# singing bird model
notes | RC == 0 ~ poisson(lambda),
notes | RC == 1 ~ custom(log_sum_exp(
log(k) + poisson_lpmf(notes | exp(a + b)),
log(1 - k) + poisson_lpmf(notes | exp(a))
)),
log(lambda) <- a + b * cat,
a ~ normal(5, 3),
b ~ normal(-3, 2),
# sneaking cat model
cat | RC == 0 ~ bernoulli(k),
k ~ beta(2, 2),
# imputed values
gq > vector[N]:PrC1 <- exp(lpC1) / (exp(lpC1) + exp(lpC0)),
gq > vector[N]:lpC1 <- log(k) + poisson_lpmf(notes[i] | exp(a + b)),
gq > vector[N]:lpC0 <- log(1 - k) + poisson_lpmf(notes[i] | exp(a))
),
cmdstan = TRUE, data = dat, chains = 4, cores = 4
)
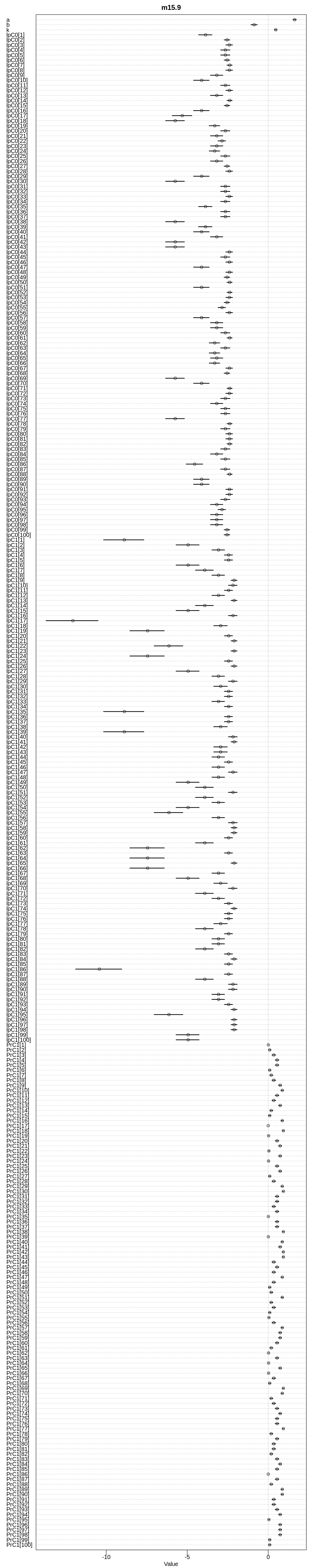
display(precis(m15.9, depth=3), mimetypes="text/plain")
iplot(function() {
plot(precis(m15.9, depth=3), main="m15.9")
}, ar=0.2)
Warning in '/tmp/RtmpjPjDJw/model-122e748371.stan', line 3, column 4: Declaration
of arrays by placing brackets after a variable name is deprecated and
will be removed in Stan 2.32.0. Instead use the array keyword before the
type. This can be changed automatically using the auto-format flag to
stanc
Warning in '/tmp/RtmpjPjDJw/model-122e748371.stan', line 4, column 4: Declaration
of arrays by placing brackets after a variable name is deprecated and
will be removed in Stan 2.32.0. Instead use the array keyword before the
type. This can be changed automatically using the auto-format flag to
stanc
Warning in '/tmp/RtmpjPjDJw/model-122e748371.stan', line 5, column 4: Declaration
of arrays by placing brackets after a variable name is deprecated and
will be removed in Stan 2.32.0. Instead use the array keyword before the
type. This can be changed automatically using the auto-format flag to
stanc
Running MCMC with 4 parallel chains, with 1 thread(s) per chain...
Chain 1 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 1 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 1 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 1 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 2 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 2 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 2 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 2 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 3 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 3 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 3 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 4 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 4 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 4 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 1 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 1 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 1 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 1 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 2 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 2 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 2 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 2 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 3 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 3 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 3 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 3 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 4 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 4 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 4 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 4 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 1 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 1 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 1 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 2 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 2 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 3 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 3 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 3 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 4 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 4 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 1 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 2 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 2 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 3 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 3 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 4 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 4 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 1 finished in 0.4 seconds.
Chain 2 finished in 0.4 seconds.
Chain 3 finished in 0.4 seconds.
Chain 4 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 4 finished in 0.5 seconds.
All 4 chains finished successfully.
Mean chain execution time: 0.4 seconds.
Total execution time: 0.6 seconds.
mean sd 5.5% 94.5% n_eff Rhat4
a 1.6305739 0.06487360 1.5270624 1.7341393 1246.262 1.0007234
b -0.8686414 0.12404909 -1.0655309 -0.6745582 1370.308 1.0002001
k 0.4613826 0.05293322 0.3792623 0.5478094 1578.813 0.9998864
lpC0[1] -3.8678825 0.25926699 -4.2997987 -3.4731018 1307.226 1.0008298
lpC0[2] -2.5370281 0.10940782 -2.7196871 -2.3717162 1604.795 1.0003821
lpC0[3] -2.3969789 0.13123678 -2.6149412 -2.2036879 1472.008 1.0000513
lpC0[4] -2.6412583 0.18020927 -2.9358888 -2.3658676 1357.050 1.0002515
lpC0[5] -2.6412583 0.18020927 -2.9358888 -2.3658676 1357.050 1.0002515
lpC0[6] -2.5370281 0.10940782 -2.7196871 -2.3717162 1604.795 1.0003821
lpC0[7] -2.3758426 0.10192148 -2.5441358 -2.2288279 1564.462 0.9999698
lpC0[8] -2.3969789 0.13123678 -2.6149412 -2.2036879 1472.008 1.0000513
lpC0[9] -3.1732201 0.23694897 -3.5641388 -2.8074180 1306.062 1.0003836
lpC0[10] -4.1106469 0.29703792 -4.6066845 -3.6474712 1282.459 1.0004647
lpC0[11] -2.6412583 0.18020927 -2.9358888 -2.3658676 1357.050 1.0002515
lpC0[12] -2.3969789 0.13123678 -2.6149412 -2.2036879 1472.008 1.0000513
lpC0[13] -3.1732201 0.23694897 -3.5641388 -2.8074180 1306.062 1.0003836
lpC0[14] -2.3758426 0.10192148 -2.5441358 -2.2288279 1564.462 0.9999698
lpC0[15] -2.5370281 0.10940782 -2.7196871 -2.3717162 1604.795 1.0003821
lpC0[16] -4.1106469 0.29703792 -4.6066845 -3.6474712 1282.459 1.0004647
lpC0[17] -5.3072147 0.38228011 -5.9249241 -4.7161928 1275.016 1.0008237
lpC0[18] -5.7412210 0.35879703 -6.3358815 -5.1745468 1270.034 1.0005170
lpC0[19] -3.3012319 0.20096205 -3.6413410 -2.9936611 1348.218 1.0008078
lpC0[20] -2.6412583 0.18020927 -2.9358888 -2.3658676 1357.050 1.0002515
lpC0[21] -3.1732201 0.23694897 -3.5641388 -2.8074180 1306.062 1.0003836
lpC0[22] -2.8523643 0.14822054 -3.0976313 -2.6325850 1441.347 1.0007080
lpC0[23] -3.1732201 0.23694897 -3.5641388 -2.8074180 1306.062 1.0003836
lpC0[24] -3.3012319 0.20096205 -3.6413410 -2.9936611 1348.218 1.0008078
lpC0[25] -2.6412583 0.18020927 -2.9358888 -2.3658676 1357.050 1.0002515
lpC0[26] -3.1732201 0.23694897 -3.5641388 -2.8074180 1306.062 1.0003836
lpC0[27] -2.5370281 0.10940782 -2.7196871 -2.3717162 1604.795 1.0003821
⋮ ⋮ ⋮ ⋮ ⋮ ⋮ ⋮
PrC1[71] 0.184080542 0.057826721 0.1004002350 0.28357880 1680.333 0.9999298
PrC1[72] 0.342496963 0.071722656 0.2327824950 0.45816177 1658.033 0.9997594
PrC1[73] 0.548759528 0.071358327 0.4348512500 0.65723994 1568.274 0.9998947
PrC1[74] 0.739902231 0.058983217 0.6397687550 0.82782831 1374.728 1.0002978
PrC1[75] 0.548759528 0.071358327 0.4348512500 0.65723994 1568.274 0.9998947
PrC1[76] 0.548759528 0.071358327 0.4348512500 0.65723994 1568.274 0.9998947
PrC1[77] 0.937878873 0.027476536 0.8864772500 0.97174892 1068.440 1.0006343
PrC1[78] 0.184080542 0.057826721 0.1004002350 0.28357880 1680.333 0.9999298
PrC1[79] 0.548759528 0.071358327 0.4348512500 0.65723994 1568.274 0.9998947
PrC1[80] 0.342496963 0.071722656 0.2327824950 0.45816177 1658.033 0.9997594
PrC1[81] 0.342496963 0.071722656 0.2327824950 0.45816177 1658.033 0.9997594
PrC1[82] 0.184080542 0.057826721 0.1004002350 0.28357880 1680.333 0.9999298
PrC1[83] 0.548759528 0.071358327 0.4348512500 0.65723994 1568.274 0.9998947
PrC1[84] 0.739902231 0.058983217 0.6397687550 0.82782831 1374.728 1.0002978
PrC1[85] 0.548759528 0.071358327 0.4348512500 0.65723994 1568.274 0.9998947
PrC1[86] 0.004186114 0.004196283 0.0006075278 0.01178349 1601.748 1.0012753
PrC1[87] 0.548759528 0.071358327 0.4348512500 0.65723994 1568.274 0.9998947
PrC1[88] 0.184080542 0.057826721 0.1004002350 0.28357880 1680.333 0.9999298
PrC1[89] 0.868392604 0.042487911 0.7907626650 0.92642500 1154.959 1.0005724
PrC1[90] 0.868392604 0.042487911 0.7907626650 0.92642500 1154.959 1.0005724
PrC1[91] 0.342496963 0.071722656 0.2327824950 0.45816177 1658.033 0.9997594
PrC1[92] 0.342496963 0.071722656 0.2327824950 0.45816177 1658.033 0.9997594
PrC1[93] 0.548759528 0.071358327 0.4348512500 0.65723994 1568.274 0.9998947
PrC1[94] 0.739902231 0.058983217 0.6397687550 0.82782831 1374.728 1.0002978
PrC1[95] 0.042253297 0.023729624 0.0141282785 0.08730985 1678.285 1.0005301
PrC1[96] 0.739902231 0.058983217 0.6397687550 0.82782831 1374.728 1.0002978
PrC1[97] 0.739902231 0.058983217 0.6397687550 0.82782831 1374.728 1.0002978
PrC1[98] 0.739902231 0.058983217 0.6397687550 0.82782831 1374.728 1.0002978
PrC1[99] 0.090158074 0.039075072 0.0389671790 0.16067187 1679.383 1.0002151
PrC1[100] 0.090158074 0.039075072 0.0389671790 0.16067187 1679.383 1.0002151
Indexes of the missing values:
display(which(R_C != 0))
post <- extract.samples(m15.9)
- 1
- 4
- 14
- 22
- 26
- 28
- 29
- 30
- 32
- 41
- 46
- 47
- 48
- 53
- 55
- 59
- 60
- 65
- 66
- 81
- 83
- 84
- 92
- 100
The mean values of PrC1 at those indexes:
mean_prc1 <- apply(post$PrC1, 2, mean)
display(mean_prc1[R_C != 0])
- 0.00900593709
- 0.5487595275
- 0.1840805422
- 0.04225329709
- 0.7399022305
- 0.3424969625
- 0.8683926035
- 0.9378788725
- 0.5487595275
- 0.7399022305
- 0.3424969625
- 0.8683926035
- 0.3424969625
- 0.3424969625
- 0.04225329709
- 0.7399022305
- 0.5487595275
- 0.7399022305
- 0.01950867139
- 0.3424969625
- 0.5487595275
- 0.7399022305
- 0.3424969625
- 0.0901580741
The true simulated values at those indexes:
display(cat[R_C != 0])
- 0
- 0
- 0
- 0
- 1
- 0
- 1
- 1
- 1
- 1
- 0
- 1
- 1
- 1
- 0
- 1
- 0
- 1
- 0
- 0
- 0
- 1
- 0
- 0
If we take PrC1 > 0.5 to mean a cat is present, then these missing elements were predicted
correctly:
cat_prc1 <- as.integer(mean_prc1 > 0.5)
is_correct <- cat_prc1[R_C != 0] == cat[R_C != 0]
display(is_correct)
- TRUE
- FALSE
- TRUE
- TRUE
- TRUE
- TRUE
- TRUE
- TRUE
- TRUE
- TRUE
- TRUE
- TRUE
- FALSE
- FALSE
- TRUE
- TRUE
- FALSE
- TRUE
- TRUE
- TRUE
- FALSE
- TRUE
- TRUE
- TRUE
For a total accuracy of:
display(sum(is_correct) / length(is_correct))
15M6. Return to the four dog-eats-homework missing data examples. Simulate each and then fit one or more models to try to recover valid estimates for \(S \rightarrow H\).
Answer. A model fit to completely observed (co) data and a model fit to complete cases (cc)
for the first scenario, where dogs eat homework at random:
library(data.table)
N <- 100
S <- rnorm(N)
H <- rbinom(N, size = 10, inv_logit(S))
D <- rbern(N) # dogs completely random
Hm <- H
Hm[D == 1] <- NA
d <- data.frame(S = S, H = H, Hm = Hm)
check_co <- function(d, name) {
set.seed(27)
dat.co <- list(
S = d$S,
H = d$H
)
m.dog.co <- ulam(
alist(
H ~ dbinom(10, p),
logit(p) <- a + bS*S,
a ~ dnorm(0, 1),
bS ~ dnorm(0, 1)
), cmdstan = TRUE, data=dat.co, cores=4, chains=4
)
display(precis(m.dog.co, depth=2), mimetypes="text/plain")
iplot(function() {
plot(precis(m.dog.co, depth=2), main=name)
}, ar=4.0)
}
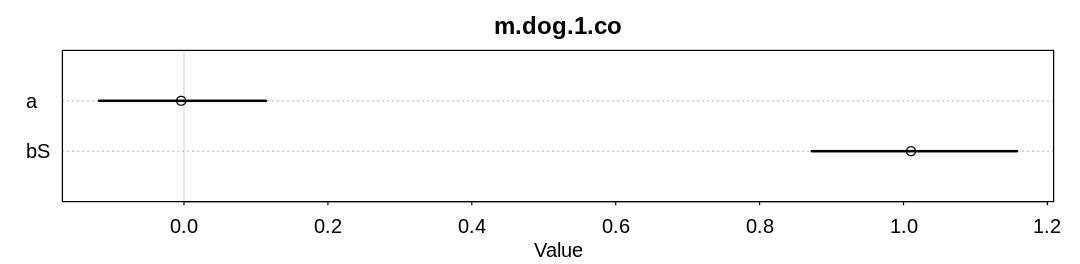
check_co(d, "m.dog.1.co")
check_cc <- function(d, name) {
# browser()
set.seed(27)
d_cc <- d[complete.cases(d$Hm), ]
# t_d <- transpose(d_cc)
# colnames(t_d) <- rownames(d_cc)
# rownames(t_d) <- colnames(d_cc)
# display(t_d)
dat.cc <- list(
S = d_cc$S,
Hm = d_cc$Hm
)
display_markdown("Sample of data remaining for 'complete case' analysis: ")
display(head(d_cc))
m.dog.cc <- ulam(
alist(
Hm ~ dbinom(10, p),
logit(p) <- a + bS*S,
a ~ dnorm(0, 1),
bS ~ dnorm(0, 1)
), cmdstan = TRUE, data=dat.cc, cores=4, chains=4
)
display(precis(m.dog.cc, depth=2), mimetypes="text/plain")
iplot(function() {
plot(precis(m.dog.cc, depth=2), main=name)
}, ar=4.0)
}
check_cc(d, "m.dog.1.cc")
Warning in '/tmp/RtmpjPjDJw/model-1215642add.stan', line 2, column 4: Declaration
of arrays by placing brackets after a variable name is deprecated and
will be removed in Stan 2.32.0. Instead use the array keyword before the
type. This can be changed automatically using the auto-format flag to
stanc
mean sd 5.5% 94.5% n_eff Rhat4
a -0.00386812 0.07272701 -0.1179536 0.1137859 1440.388 0.9994012
bS 1.01038932 0.08952928 0.8723994 1.1575217 1035.581 1.0002041
Running MCMC with 4 parallel chains, with 1 thread(s) per chain...
Chain 1 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 1 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 1 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 1 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 1 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 1 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 1 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 1 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 1 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 1 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 1 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 1 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 2 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 2 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 2 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 2 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 2 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 2 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 2 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 2 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 2 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 2 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 2 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 2 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 3 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 3 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 3 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 3 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 3 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 3 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 3 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 3 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 3 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 3 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 3 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 3 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 4 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 4 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 4 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 4 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 4 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 4 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 4 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 4 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 4 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 4 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 4 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 4 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 1 finished in 0.1 seconds.
Chain 2 finished in 0.1 seconds.
Chain 3 finished in 0.1 seconds.
Chain 4 finished in 0.1 seconds.
All 4 chains finished successfully.
Mean chain execution time: 0.1 seconds.
Total execution time: 0.2 seconds.
Sample of data remaining for ‘complete case’ analysis:
| S | H | Hm | |
|---|---|---|---|
| <dbl> | <int> | <int> | |
| 3 | -0.3483160 | 6 | 6 |
| 5 | 0.2841523 | 5 | 5 |
| 8 | 0.3812747 | 5 | 5 |
| 10 | 0.3922908 | 8 | 8 |
| 13 | 0.1487904 | 8 | 8 |
| 16 | 1.1348391 | 9 | 9 |
Warning in '/tmp/RtmpjPjDJw/model-1230488a32.stan', line 2, column 4: Declaration
of arrays by placing brackets after a variable name is deprecated and
will be removed in Stan 2.32.0. Instead use the array keyword before the
type. This can be changed automatically using the auto-format flag to
stanc
mean sd 5.5% 94.5% n_eff Rhat4
a -0.03654073 0.09791599 -0.1900768 0.1254365 1339.579 1.001207
bS 0.92498400 0.12369701 0.7329099 1.1265327 1116.149 1.000748
Running MCMC with 4 parallel chains, with 1 thread(s) per chain...
Chain 1 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 1 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 1 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 1 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 1 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 1 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 1 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 1 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 1 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 1 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 1 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 1 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 2 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 2 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 2 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 2 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 2 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 2 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 2 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 2 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 2 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 2 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 2 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 2 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 3 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 3 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 3 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 3 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 3 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 3 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 3 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 3 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 3 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 3 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 3 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 3 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 4 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 4 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 4 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 4 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 4 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 4 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 4 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 4 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 4 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 4 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 4 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 4 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 1 finished in 0.1 seconds.
Chain 2 finished in 0.1 seconds.
Chain 3 finished in 0.1 seconds.
Chain 4 finished in 0.1 seconds.
All 4 chains finished successfully.
Mean chain execution time: 0.1 seconds.
Total execution time: 0.2 seconds.
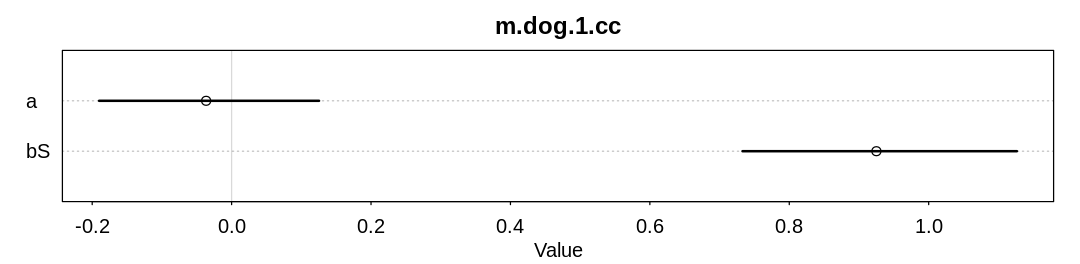

As expected (explained in the chapter) we are able to infer bS both with and without all the data.
In the second scenario, dogs of hard-working students eat homework. We don’t need to fit a new
completely observed (co) model because it shouldn’t be any different from m.dog.1.co. A model
fit to complete cases (cc):
D2 <- ifelse(S > 0, 1, 0)
Hm2 <- H
Hm2[D2 == 1] <- NA
d <- data.frame(S = S, H = H, Hm = Hm2)
check_cc(d, "m.dog.2.cc")
Sample of data remaining for ‘complete case’ analysis:
| S | H | Hm | |
|---|---|---|---|
| <dbl> | <int> | <int> | |
| 2 | -1.29318475 | 2 | 2 |
| 3 | -0.34831602 | 6 | 6 |
| 4 | -0.35097548 | 2 | 2 |
| 20 | -1.44522679 | 2 | 2 |
| 21 | -0.68131726 | 2 | 2 |
| 24 | -0.04805382 | 6 | 6 |
Warning in '/tmp/RtmpjPjDJw/model-1247d6a9ed.stan', line 2, column 4: Declaration
of arrays by placing brackets after a variable name is deprecated and
will be removed in Stan 2.32.0. Instead use the array keyword before the
type. This can be changed automatically using the auto-format flag to
stanc
mean sd 5.5% 94.5% n_eff Rhat4
a -0.2211787 0.1625756 -0.4830884 0.03894572 567.4133 1.007038
bS 0.8135978 0.2028660 0.4879648 1.14727990 421.4382 1.008410
Running MCMC with 4 parallel chains, with 1 thread(s) per chain...
Chain 1 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 1 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 1 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 1 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 1 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 1 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 1 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 1 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 1 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 1 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 1 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 1 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 2 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 2 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 2 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 2 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 2 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 2 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 2 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 2 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 2 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 2 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 2 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 2 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 3 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 3 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 3 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 3 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 3 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 3 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 3 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 3 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 3 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 3 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 3 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 3 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 4 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 4 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 4 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 4 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 4 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 4 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 4 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 4 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 4 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 4 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 4 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 4 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 1 finished in 0.1 seconds.
Chain 2 finished in 0.1 seconds.
Chain 3 finished in 0.1 seconds.
Chain 4 finished in 0.1 seconds.
All 4 chains finished successfully.
Mean chain execution time: 0.1 seconds.
Total execution time: 0.2 seconds.
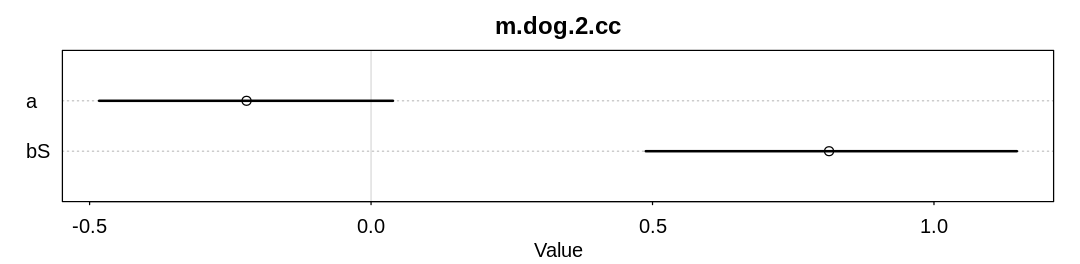
As expected (explained in the chapter) we are able to infer bS both with and without all the data.
The text suggests a variation on scenario 2 where the function \(S \rightarrow H\) is nonlinear and unobserved only in the domain of the function where it is non-linear. Let’s simulate this scenario and attempt to infer the coefficient of the (wrong) linear function with and without all the data:
H3 <- ifelse(S > 0, H, 0)
Hm3 <- H3
Hm3[D2 == 1] <- NA
d <- data.frame(S = S, H = H3, Hm = Hm3)
check_co(d, "m.dog.2.var.co")
check_cc(d, "m.dog.2.var.cc")
Warning in '/tmp/RtmpjPjDJw/model-127154fd95.stan', line 2, column 4: Declaration
of arrays by placing brackets after a variable name is deprecated and
will be removed in Stan 2.32.0. Instead use the array keyword before the
type. This can be changed automatically using the auto-format flag to
stanc
mean sd 5.5% 94.5% n_eff Rhat4
a -1.065430 0.09883981 -1.225324 -0.9143453 983.0190 1.001233
bS 2.289743 0.14768992 2.061293 2.5359345 849.6928 1.001852
Running MCMC with 4 parallel chains, with 1 thread(s) per chain...
Chain 1 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 1 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 1 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 1 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 1 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 1 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 1 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 1 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 1 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 1 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 1 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 1 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 2 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 2 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 2 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 2 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 2 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 2 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 2 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 2 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 2 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 2 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 2 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 2 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 3 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 3 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 3 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 3 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 3 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 3 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 3 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 3 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 3 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 3 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 3 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 3 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 4 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 4 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 4 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 4 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 4 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 4 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 4 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 4 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 4 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 4 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 4 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 4 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 1 finished in 0.1 seconds.
Chain 2 finished in 0.1 seconds.
Chain 3 finished in 0.1 seconds.
Chain 4 finished in 0.1 seconds.
All 4 chains finished successfully.
Mean chain execution time: 0.1 seconds.
Total execution time: 0.2 seconds.
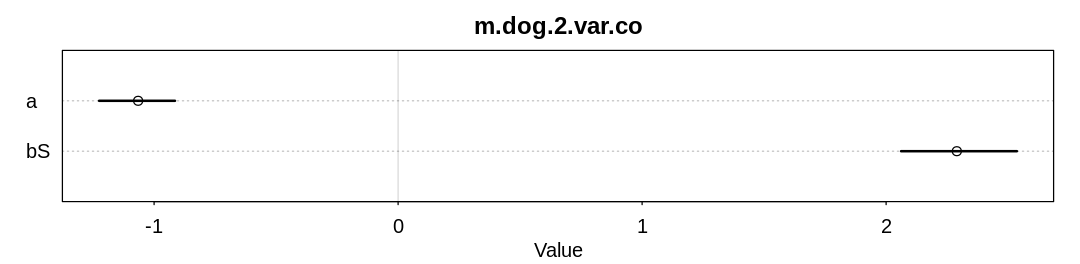
Sample of data remaining for ‘complete case’ analysis:
| S | H | Hm | |
|---|---|---|---|
| <dbl> | <dbl> | <dbl> | |
| 2 | -1.29318475 | 0 | 0 |
| 3 | -0.34831602 | 0 | 0 |
| 4 | -0.35097548 | 0 | 0 |
| 20 | -1.44522679 | 0 | 0 |
| 21 | -0.68131726 | 0 | 0 |
| 24 | -0.04805382 | 0 | 0 |
Warning in '/tmp/RtmpjPjDJw/model-1222d838f9.stan', line 2, column 4: Declaration
of arrays by placing brackets after a variable name is deprecated and
will be removed in Stan 2.32.0. Instead use the array keyword before the
type. This can be changed automatically using the auto-format flag to
stanc
mean sd 5.5% 94.5% n_eff Rhat4
a -3.937813 0.4984334 -4.7425330 -3.171735 937.6243 1.003867
bS 1.580369 0.7318090 0.4746245 2.783566 943.3718 1.004776
Running MCMC with 4 parallel chains, with 1 thread(s) per chain...
Chain 1 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 1 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 1 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 1 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 1 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 1 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 1 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 1 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 1 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 1 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 1 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 1 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 2 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 2 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 2 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 2 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 2 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 2 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 2 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 2 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 2 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 2 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 2 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 2 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 3 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 3 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 3 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 3 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 3 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 3 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 3 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 3 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 3 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 3 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 3 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 3 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 4 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 4 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 4 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 4 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 4 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 4 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 4 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 4 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 4 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 4 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 4 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 4 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 1 finished in 0.1 seconds.
Chain 2 finished in 0.1 seconds.
Chain 3 finished in 0.1 seconds.
Chain 4 finished in 0.1 seconds.
All 4 chains finished successfully.
Mean chain execution time: 0.1 seconds.
Total execution time: 0.2 seconds.
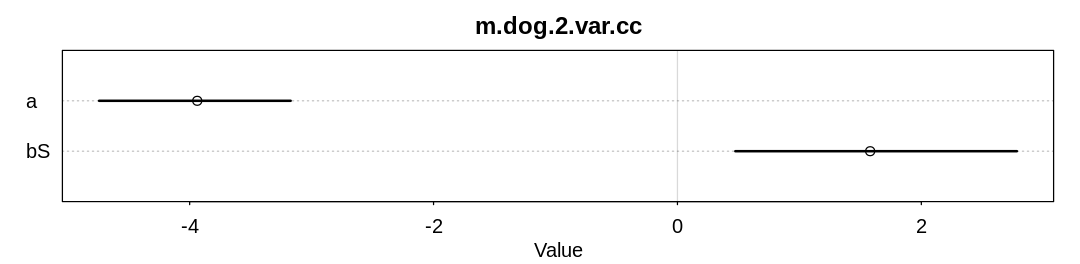
As expected (explained in the chapter) both these inferences are wrong. The cc inference is wrong
because despite complete observability our model was linear when the function was non-linear. The
co inference is extra wrong because our sampling is also biased. The data going into this second
model is completely uninformative; all the S are random negative numbers and the H are 0. Notice
that we confusingly still get a positive inference for bS. The model wants to push everything in
the log link as negative as possible, but a is already as negative as it can be given our thin
tailed (normal) prior on it. We can push everything farther negative by making bS positive, since
all the S values are negative. Be skeptical of models that have any parameter inferences that are
extreme relative to the parameter’s priors (like a)! These mean that either your prior is wrong,
your data, or your model.
The third scenario, where some unobserved variable like the noise level in the student’s house affects both the homework quality and whether dogs eat homework, is already partially covered in the chapter. Let’s consider the variation suggested in R code 15.14, however:
N2 <- 1000
X <- rnorm(N2)
S2 <- rnorm(N2)
H4 <- rbinom(N2, size = 10, inv_logit(2 + S2 - 2 * X))
# D <- ifelse(X > 1, 1, 0)
D <- ifelse(abs(X) < 1, 1, 0)
Hm4 <- H4
Hm4[D == 1] <- NA
d <- data.frame(S = S2, H = H4, Hm = Hm4)
check_co(d, "m.dog.3.var.co")
check_cc(d, "m.dog.3.var.cc")
Warning in '/tmp/RtmpjPjDJw/model-122069a9e2.stan', line 2, column 4: Declaration
of arrays by placing brackets after a variable name is deprecated and
will be removed in Stan 2.32.0. Instead use the array keyword before the
type. This can be changed automatically using the auto-format flag to
stanc
mean sd 5.5% 94.5% n_eff Rhat4
a 1.2221746 0.02611138 1.1807394 1.2637615 1019.2430 0.9993861
bS 0.6320652 0.02500255 0.5934007 0.6724795 944.8489 1.0001442
Running MCMC with 4 parallel chains, with 1 thread(s) per chain...
Chain 1 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 1 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 2 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 2 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 3 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 3 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 4 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 4 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 1 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 1 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 2 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 2 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 3 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 3 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 4 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 1 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 1 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 1 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 2 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 3 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 4 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 1 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 2 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 2 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 3 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 3 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 3 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 4 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 4 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 4 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 1 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 1 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 2 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 2 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 3 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 4 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 1 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 2 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 3 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 4 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 1 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 2 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 2 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 3 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 4 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 4 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 1 finished in 0.7 seconds.
Chain 2 finished in 0.8 seconds.
Chain 3 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 4 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 3 finished in 0.8 seconds.
Chain 4 finished in 0.8 seconds.
All 4 chains finished successfully.
Mean chain execution time: 0.8 seconds.
Total execution time: 0.9 seconds.
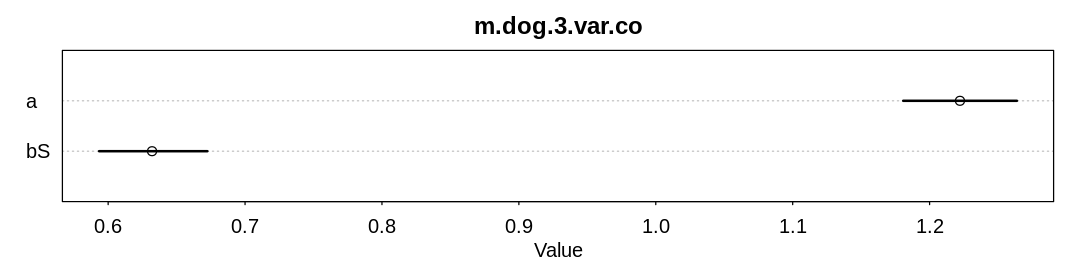
Sample of data remaining for ‘complete case’ analysis:
| S | H | Hm | |
|---|---|---|---|
| <dbl> | <int> | <int> | |
| 1 | 0.91583404 | 3 | 3 |
| 3 | -0.75727948 | 9 | 9 |
| 8 | 0.06453134 | 10 | 10 |
| 9 | 1.22364898 | 10 | 10 |
| 11 | -0.85843194 | 10 | 10 |
| 14 | 0.12535586 | 10 | 10 |
Warning in '/tmp/RtmpjPjDJw/model-121196a3b8.stan', line 2, column 4: Declaration
of arrays by placing brackets after a variable name is deprecated and
will be removed in Stan 2.32.0. Instead use the array keyword before the
type. This can be changed automatically using the auto-format flag to
stanc
mean sd 5.5% 94.5% n_eff Rhat4
a 0.6269566 0.03576682 0.5709384 0.6832952 1348.5960 1.000598
bS 0.3857848 0.03922876 0.3241633 0.4471760 732.3931 1.005366
Running MCMC with 4 parallel chains, with 1 thread(s) per chain...
Chain 1 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 1 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 1 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 1 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 1 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 1 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 1 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 2 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 2 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 2 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 2 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 2 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 2 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 2 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 3 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 3 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 3 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 3 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 3 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 3 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 3 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 4 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 4 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 4 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 4 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 4 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 1 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 1 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 1 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 1 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 1 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 2 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 2 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 2 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 2 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 2 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 3 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 3 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 3 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 3 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 4 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 4 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 4 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 4 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 4 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 1 finished in 0.2 seconds.
Chain 2 finished in 0.3 seconds.
Chain 3 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 4 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 4 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 3 finished in 0.3 seconds.
Chain 4 finished in 0.3 seconds.
All 4 chains finished successfully.
Mean chain execution time: 0.3 seconds.
Total execution time: 0.4 seconds.
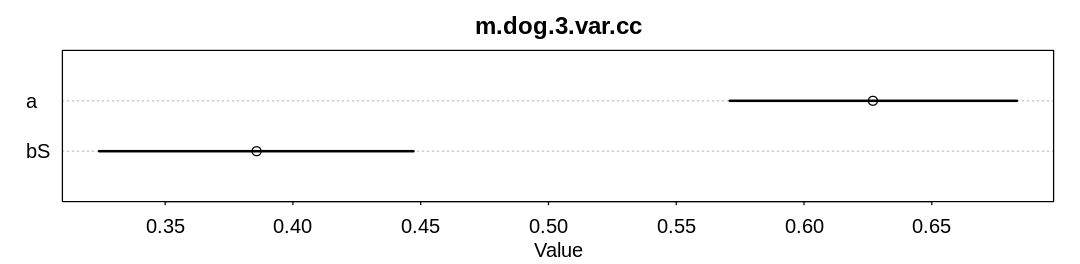
As expected (see the text) missingness makes the inference for bS worse than with complete
observability. We’ve changed the missingness function to remove small values rather than large
values. When we remove large values, the remaining observations are farther from a ceiling and
therefore the influence of predictors seems larger, which is what we want when we have an unobserved
variable. When we remove small values, the remaining observations are closer to a ceiling or a
floor and the influence of predictors seems smaller.
In the fourth scenario dogs eat bad homework. Little can be done to get a good inference:
S.4 <- rnorm(N)
H.4 <- rbinom(N, size = 10, inv_logit(S.4))
D.4 <- ifelse(H.4 < 5, 1, 0)
Hm.4 <- H.4
Hm.4[D.4 == 1] <- NA
d <- data.frame(S = S.4, H = H.4, Hm = Hm.4)
check_cc(d, "m.dog.4.cc")
Sample of data remaining for ‘complete case’ analysis:
| S | H | Hm | |
|---|---|---|---|
| <dbl> | <int> | <int> | |
| 1 | 1.67531074 | 8 | 8 |
| 2 | 0.80350151 | 7 | 7 |
| 5 | 0.25107932 | 7 | 7 |
| 7 | 0.35701984 | 6 | 6 |
| 12 | 0.05818869 | 5 | 5 |
| 16 | 0.18096825 | 5 | 5 |
Warning in '/tmp/RtmpjPjDJw/model-123483df95.stan', line 2, column 4: Declaration
of arrays by placing brackets after a variable name is deprecated and
will be removed in Stan 2.32.0. Instead use the array keyword before the
type. This can be changed automatically using the auto-format flag to
stanc
mean sd 5.5% 94.5% n_eff Rhat4
a 0.3559647 0.1110200 0.1814343 0.5382437 702.4761 0.9993733
bS 0.6936059 0.1352675 0.4730826 0.9099878 747.4805 1.0009603
Running MCMC with 4 parallel chains, with 1 thread(s) per chain...
Chain 1 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 1 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 1 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 1 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 1 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 1 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 1 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 1 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 1 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 1 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 1 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 1 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 2 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 2 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 2 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 2 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 2 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 2 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 2 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 2 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 2 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 2 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 2 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 2 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 3 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 3 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 3 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 3 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 3 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 3 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 3 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 3 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 3 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 3 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 3 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 3 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 4 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 4 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 4 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 4 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 4 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 4 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 4 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 4 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 4 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 4 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 4 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 4 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 1 finished in 0.1 seconds.
Chain 2 finished in 0.1 seconds.
Chain 3 finished in 0.1 seconds.
Chain 4 finished in 0.1 seconds.
All 4 chains finished successfully.
Mean chain execution time: 0.1 seconds.
Total execution time: 0.2 seconds.
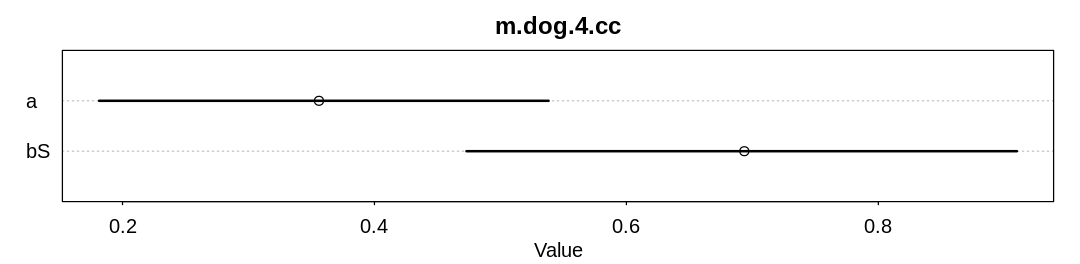
15H1. The data in data(elephants) are counts of matings observed for bull elephants of
differing ages. There is a strong positive relationship between age and matings. However, age is not
always assessed accurately. First, fit a Poisson model predicting MATINGS with AGE as a
predictor. Second, assume that the observed AGE values are uncertain and have a standard error of
±5 years. Re-estimate the relationship between MATINGS and AGE, incorporating this measurement
error. Compare the inferences of the two models.
Answer. The full elephants data.frame:
data(elephants)
display(elephants)
q15.h1.a <- function(d) {
dat <- list(
M = as.integer(d$MATINGS),
A = as.numeric(standardize(d$AGE))
)
m15.h1.a <- ulam(
alist(
M ~ dpois(lambda),
lambda <- exp(a + bA*A),
a ~ dnorm(0, 1),
bA ~ dnorm(0, 1)
), cmdstan = TRUE, data=dat, cores = 4, chains = 4, log_lik = TRUE
)
display_precis(m15.h1.a, "m15.h1.a", 4.0)
return(m15.h1.a)
}
q15.h1.b <- function(d, error=5.0) {
set.seed(31)
stdA = standardize(d$AGE)
dat <- list(
M = as.integer(d$MATINGS),
A_obs = as.numeric(stdA),
A_sd = error / attr(stdA, "scaled:scale"),
N = nrow(d)
)
m15.h1.b <- ulam(
alist(
M ~ dpois(lambda),
A_obs ~ dnorm(A_true, A_sd),
lambda <- exp(a + bA*A_true[i]),
vector[N]:A_true ~ dnorm(0, 1),
a ~ dnorm(0, 1),
bA ~ dnorm(0, 1)
), cmdstan = TRUE, data=dat, cores = 4, chains = 4, log_lik = TRUE
)
display_precis(m15.h1.b, "m15.h1.b", 0.8)
return(m15.h1.b)
}
# m15.h1.a <- q15.h1.a(elephants)
# m15.h1.b <- q15.h1.b(elephants)
#
# iplot(function() {
# plot(coeftab(m15.h1.b, m15.h1.a), pars=c("a", "bA"), main="Parameter comparison for elephant models")
# }, ar=3.0)
| AGE | MATINGS |
|---|---|
| <int> | <int> |
| 27 | 0 |
| 28 | 1 |
| 28 | 1 |
| 28 | 1 |
| 28 | 3 |
| 29 | 0 |
| 29 | 0 |
| 29 | 0 |
| 29 | 2 |
| 29 | 2 |
| 29 | 2 |
| 30 | 1 |
| 32 | 2 |
| 33 | 4 |
| 33 | 3 |
| 33 | 3 |
| 33 | 3 |
| 33 | 2 |
| 34 | 1 |
| 34 | 1 |
| 34 | 2 |
| 34 | 3 |
| 36 | 5 |
| 36 | 6 |
| 37 | 1 |
| 37 | 1 |
| 37 | 6 |
| 38 | 2 |
| 39 | 1 |
| 41 | 3 |
| 42 | 4 |
| 43 | 0 |
| 43 | 2 |
| 43 | 3 |
| 43 | 4 |
| 43 | 9 |
| 44 | 3 |
| 45 | 5 |
| 47 | 7 |
| 48 | 2 |
| 52 | 9 |
The second model infers both a larger and more uncertain estimate (wider credible intervals) for
bA.
15H2. Repeat the model fitting problem above, now increasing the assumed standard error on
AGE. How large does the standard error have to get before the posterior mean for the coefficient
on AGE reaches zero?
Answer. The coefficient’s uncertainty gets large enough to cover zero when we get to roughly 20-25 years of age uncertainty:
q15.h1.b(elephants, error=25)
Warning in '/tmp/RtmpjPjDJw/model-12743d4de1.stan', line 3, column 4: Declaration
of arrays by placing brackets after a variable name is deprecated and
will be removed in Stan 2.32.0. Instead use the array keyword before the
type. This can be changed automatically using the auto-format flag to
stanc
Running MCMC with 4 parallel chains, with 1 thread(s) per chain...
Chain 1 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 1 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 1 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 1 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 2 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 2 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 2 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 2 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 3 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 3 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 3 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 3 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 4 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 4 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 4 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 1 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 1 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 1 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 1 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 1 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 1 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 1 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 1 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 2 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 2 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 2 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 2 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 2 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 2 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 2 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 2 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 3 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 3 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 3 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 3 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 3 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 3 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 3 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 4 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 4 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 4 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 4 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 4 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 4 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 4 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 1 finished in 0.3 seconds.
Chain 2 finished in 0.3 seconds.
Chain 3 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 4 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 4 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 3 finished in 0.3 seconds.
Chain 4 finished in 0.3 seconds.
All 4 chains finished successfully.
Mean chain execution time: 0.3 seconds.
Total execution time: 0.4 seconds.
Raw data (following plot):
mean sd 5.5% 94.5% n_eff Rhat4
A_true[1] -0.60117469 1.0091844 -2.0722995 1.0873623 14.344321 1.120206
A_true[2] -0.36787650 0.8513137 -1.7027519 0.9999452 28.260337 1.056953
A_true[3] -0.36570746 0.8614791 -1.7286127 0.9951490 33.241592 1.057124
A_true[4] -0.37982283 0.8580098 -1.7140640 0.9987858 35.989888 1.050788
A_true[5] 0.07493666 0.6996594 -1.0779699 1.1663960 595.483364 1.013497
A_true[6] -0.60846831 1.0250175 -2.1094692 1.1398493 11.788646 1.131486
A_true[7] -0.61409542 1.0013380 -2.0913275 1.0568451 12.110301 1.132563
A_true[8] -0.62425165 1.0104447 -2.2182763 1.1239754 11.248228 1.138278
A_true[9] -0.13277755 0.7341117 -1.3427536 1.0298548 1617.384900 1.006307
A_true[10] -0.13387963 0.7824652 -1.4082887 1.0681719 1429.816665 1.004412
A_true[11] -0.14135320 0.7575667 -1.3215731 1.0423720 1685.314777 1.007532
A_true[12] -0.36961043 0.8986039 -1.8168895 1.0650409 37.249962 1.053382
A_true[13] -0.11639938 0.8248625 -1.4488624 1.1664826 2303.501889 1.005051
A_true[14] 0.28239312 0.7969648 -1.0607951 1.4582005 37.533339 1.057184
A_true[15] 0.07194796 0.7655739 -1.1629314 1.2292131 1136.502776 1.009605
A_true[16] 0.09594969 0.7445255 -1.1186045 1.2374490 1617.463497 1.003458
A_true[17] 0.09693792 0.7447062 -1.0956143 1.2555275 1004.926624 1.009668
A_true[18] -0.09630907 0.7648403 -1.3584912 1.1129924 1837.140523 1.005444
A_true[19] -0.32319797 0.8443859 -1.6560516 1.0874029 27.516203 1.061404
A_true[20] -0.32582686 0.8627404 -1.7263214 1.0351113 63.919385 1.037648
A_true[21] -0.08356380 0.7170041 -1.2493470 1.0399564 3002.595613 1.001265
A_true[22] 0.10043619 0.7392507 -1.1157200 1.2598396 1280.651269 1.006613
A_true[23] 0.44967315 0.8409441 -1.0400442 1.6324739 11.980876 1.146094
A_true[24] 0.62802098 0.9662659 -1.2159055 1.9125331 9.630480 1.190744
A_true[25] -0.29544474 0.8628739 -1.6330445 1.0469239 46.111393 1.048206
A_true[26] -0.29177282 0.8602039 -1.6205196 1.1353206 49.601660 1.047328
A_true[27] 0.67260803 0.9505727 -1.1801111 1.9040425 9.303567 1.193147
A_true[28] -0.08947878 0.7602801 -1.3539159 1.0541851 2115.792679 1.003276
A_true[29] -0.30100218 0.8948009 -1.7469175 1.1253371 33.215895 1.057725
A_true[30] 0.13275653 0.7520887 -1.1455423 1.2932678 1170.307847 1.008382
A_true[31] 0.34172283 0.7643616 -0.9776645 1.5050318 31.367545 1.056077
A_true[32] -0.48979044 0.9963426 -2.0226607 1.2049862 12.988025 1.126033
A_true[33] -0.05266336 0.7399035 -1.2451754 1.0873137 2050.562082 1.003732
A_true[34] 0.14068062 0.7345973 -1.0965141 1.3081742 642.349157 1.009799
A_true[35] 0.33953829 0.7762237 -0.9566545 1.5095188 29.663239 1.058023
A_true[36] 1.09639730 1.2996878 -1.7307875 2.5033904 5.481716 1.371427
A_true[37] 0.17087846 0.7288982 -1.0229196 1.2856901 1310.801033 1.007596
A_true[38] 0.53593754 0.8557067 -0.9830917 1.7093324 10.907019 1.148637
A_true[39] 0.83153739 1.0640228 -1.3882139 2.1190046 6.843605 1.274743
A_true[40] -0.02768258 0.7587293 -1.2566854 1.1798185 1898.656795 1.004493
A_true[41] 1.12761587 1.2898501 -1.7181836 2.5366333 5.789249 1.355103
a 0.81522363 0.1435477 0.5869336 1.0458022 1203.674478 1.001330
bA 0.39535337 0.4501382 -0.6236069 0.8170445 4.838490 1.464206
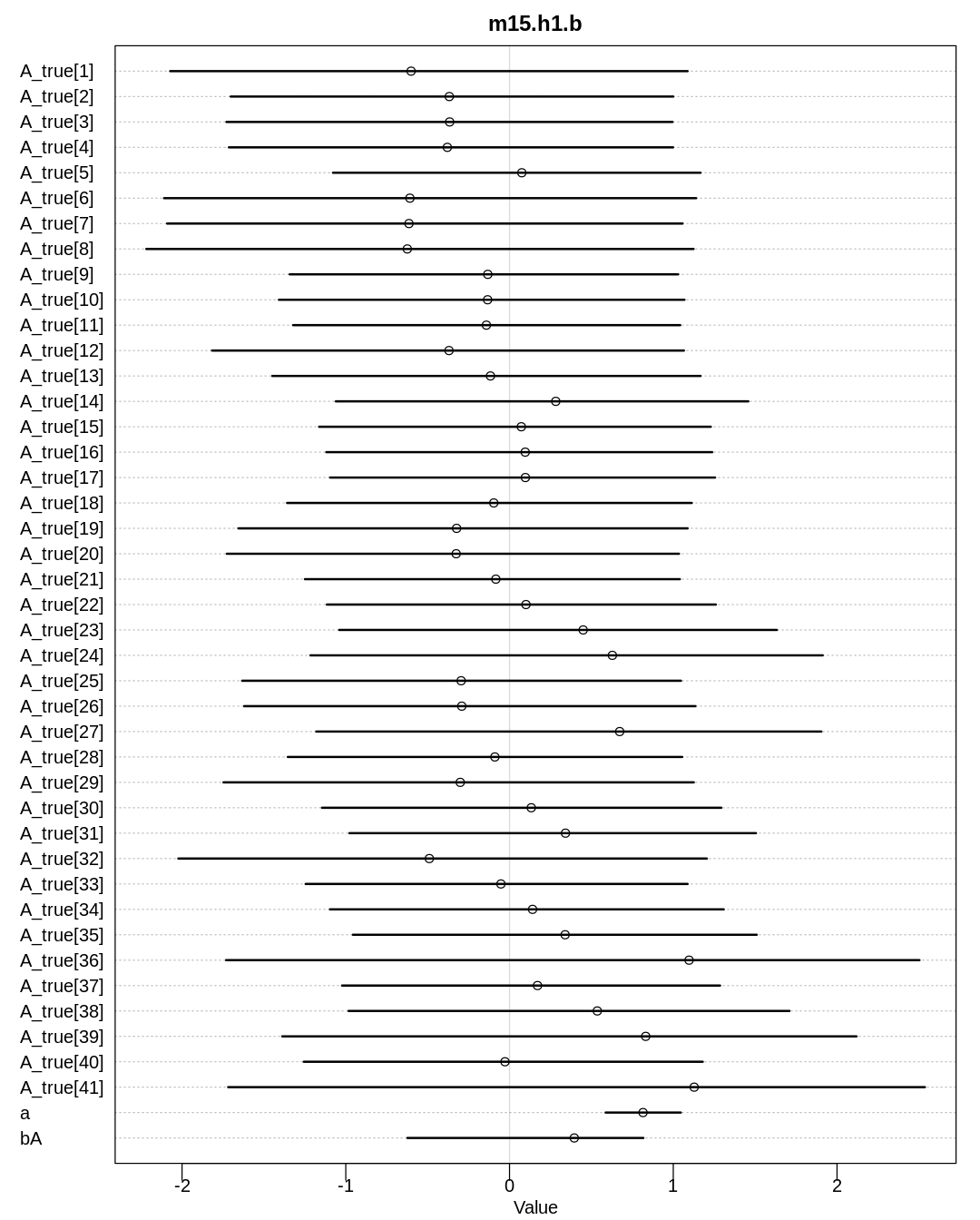
Hamiltonian Monte Carlo approximation
2000 samples from 4 chains
Sampling durations (seconds):
warmup sample total
chain:1 0.18 0.10 0.28
chain:2 0.18 0.10 0.28
chain:3 0.17 0.10 0.28
chain:4 0.18 0.11 0.29
Formula:
M ~ dpois(lambda)
A_obs ~ dnorm(A_true, A_sd)
lambda <- exp(a + bA * A_true[i])
vector[N]:A_true ~ dnorm(0, 1)
a ~ dnorm(0, 1)
bA ~ dnorm(0, 1)
15H3. The fact that information flows in all directions among parameters sometimes leads to rather unintuitive conclusions. Here’s an example from missing data imputation, in which imputation of a single datum reverses the direction of an inferred relationship. Use these data:
R code 15.32
set.seed(100)
x <- c( rnorm(10) , NA )
y <- c( rnorm(10,x) , 100 )
d <- list(x=x,y=y)
These data comprise 11 cases, one of which has a missing predictor value. You can quickly confirm
that a regression of y on x for only the complete cases indicates a strong positive relationship
between the two variables. But now fit this model, imputing the one missing value for x:
Be sure to run multiple chains. What happens to the posterior distribution of \(\beta\)? Be sure to inspect the full density. Can you explain the change in inference?
ERROR: The last line of the model should be: $\( \sigma \sim Exponential(1) \)$
Answer. The list d produced by R code 15.32:
r.code.15.32 <- function() {
set.seed(100)
x <- c(rnorm(10), NA)
y <- c(rnorm(10, x), 100)
d <- list(x = x, y = y)
display(d)
return(data.frame(d))
}
d.15.h3 <- r.code.15.32()
- $x
-
- -0.502192350531457
- 0.131531165327303
- -0.07891708981887
- 0.886784809417845
- 0.116971270510841
- 0.318630087617032
- -0.58179068471591
- 0.714532710891568
- -0.825259425862769
- -0.359862131395465
- <NA>
- $y
-
- -0.412306206753926
- 0.227805625612433
- -0.280551042002224
- 1.62662530929628
- 0.240350771599711
- 0.289313378387685
- -0.97064493161941
- 1.22538896826149
- -1.73907361123195
- 1.95043469138359
- 100
Fitting a model to complete cases, using the same model in the question minus the \(x_i\) prior:
q.15.h3.a <- function(d) {
d_cc <- d[complete.cases(d$x),]
dat <- list(
Y = d_cc$y,
X = d_cc$x
)
m15.h3.a <- ulam(
alist(
Y ~ dnorm(mu, sigma),
mu <- a + b*X,
c(a, b) ~ dnorm(0, 1),
sigma ~ dnorm(0, 1)
), cmdstan = TRUE, data=dat, cores=4, chains=4, log_lik=TRUE
)
display_precis(m15.h3.a, "m15.h3.a", 4.5)
return(m15.h3.a)
}
m15.h3.a <- q.15.h3.a(d.15.h3)
Running MCMC with 4 parallel chains, with 1 thread(s) per chain...
Chain 1 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 1 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 1 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 1 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 1 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 1 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 1 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 1 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 1 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 1 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 1 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 1 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 1 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 1 Exception: normal_lpdf: Scale parameter is -4.90991, but must be positive! (in '/tmp/RtmpjPjDJw/model-122683fea6.stan', line 18, column 4 to column 29)
Chain 1 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 1 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 1
Chain 2 Rejecting initial value:
Chain 2 Error evaluating the log probability at the initial value.
Chain 2 Exception: normal_lpdf: Scale parameter is -1.99224, but must be positive! (in '/tmp/RtmpjPjDJw/model-122683fea6.stan', line 18, column 4 to column 29)
Chain 2 Exception: normal_lpdf: Scale parameter is -1.99224, but must be positive! (in '/tmp/RtmpjPjDJw/model-122683fea6.stan', line 18, column 4 to column 29)
Chain 2 Rejecting initial value:
Chain 2 Error evaluating the log probability at the initial value.
Chain 2 Exception: normal_lpdf: Scale parameter is -0.908125, but must be positive! (in '/tmp/RtmpjPjDJw/model-122683fea6.stan', line 18, column 4 to column 29)
Chain 2 Exception: normal_lpdf: Scale parameter is -0.908125, but must be positive! (in '/tmp/RtmpjPjDJw/model-122683fea6.stan', line 18, column 4 to column 29)
Chain 2 Rejecting initial value:
Chain 2 Error evaluating the log probability at the initial value.
Chain 2 Exception: normal_lpdf: Scale parameter is -0.850954, but must be positive! (in '/tmp/RtmpjPjDJw/model-122683fea6.stan', line 18, column 4 to column 29)
Chain 2 Exception: normal_lpdf: Scale parameter is -0.850954, but must be positive! (in '/tmp/RtmpjPjDJw/model-122683fea6.stan', line 18, column 4 to column 29)
Chain 2 Rejecting initial value:
Chain 2 Error evaluating the log probability at the initial value.
Chain 2 Exception: normal_lpdf: Scale parameter is -0.806948, but must be positive! (in '/tmp/RtmpjPjDJw/model-122683fea6.stan', line 18, column 4 to column 29)
Chain 2 Exception: normal_lpdf: Scale parameter is -0.806948, but must be positive! (in '/tmp/RtmpjPjDJw/model-122683fea6.stan', line 18, column 4 to column 29)
Chain 2 Rejecting initial value:
Chain 2 Error evaluating the log probability at the initial value.
Chain 2 Exception: normal_lpdf: Scale parameter is -0.966028, but must be positive! (in '/tmp/RtmpjPjDJw/model-122683fea6.stan', line 18, column 4 to column 29)
Chain 2 Exception: normal_lpdf: Scale parameter is -0.966028, but must be positive! (in '/tmp/RtmpjPjDJw/model-122683fea6.stan', line 18, column 4 to column 29)
Chain 2 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 2 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 2 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 2 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 2 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 2 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 2 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 2 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 2 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 2 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 2 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 2 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 2 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 2 Exception: normal_lpdf: Scale parameter is -1.72548, but must be positive! (in '/tmp/RtmpjPjDJw/model-122683fea6.stan', line 18, column 4 to column 29)
Chain 2 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 2 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 2
Chain 2 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 2 Exception: normal_lpdf: Scale parameter is -0.637511, but must be positive! (in '/tmp/RtmpjPjDJw/model-122683fea6.stan', line 18, column 4 to column 29)
Chain 2 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 2 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 2
Chain 2 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 2 Exception: normal_lpdf: Scale parameter is -19.6802, but must be positive! (in '/tmp/RtmpjPjDJw/model-122683fea6.stan', line 18, column 4 to column 29)
Chain 2 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 2 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 2
Chain 2 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 2 Exception: normal_lpdf: Scale parameter is -1.36893, but must be positive! (in '/tmp/RtmpjPjDJw/model-122683fea6.stan', line 18, column 4 to column 29)
Chain 2 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 2 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 2
Chain 2 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 2 Exception: normal_lpdf: Scale parameter is -0.58501, but must be positive! (in '/tmp/RtmpjPjDJw/model-122683fea6.stan', line 18, column 4 to column 29)
Chain 2 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 2 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 2
Chain 2 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 2 Exception: normal_lpdf: Scale parameter is -3.18212, but must be positive! (in '/tmp/RtmpjPjDJw/model-122683fea6.stan', line 18, column 4 to column 29)
Chain 2 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 2 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 2
Chain 2 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 2 Exception: normal_lpdf: Scale parameter is -3.81429, but must be positive! (in '/tmp/RtmpjPjDJw/model-122683fea6.stan', line 18, column 4 to column 29)
Chain 2 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 2 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 2
Chain 3 Rejecting initial value:
Chain 3 Error evaluating the log probability at the initial value.
Chain 3 Exception: normal_lpdf: Scale parameter is -1.38994, but must be positive! (in '/tmp/RtmpjPjDJw/model-122683fea6.stan', line 18, column 4 to column 29)
Chain 3 Exception: normal_lpdf: Scale parameter is -1.38994, but must be positive! (in '/tmp/RtmpjPjDJw/model-122683fea6.stan', line 18, column 4 to column 29)
Chain 3 Rejecting initial value:
Chain 3 Error evaluating the log probability at the initial value.
Chain 3 Exception: normal_lpdf: Scale parameter is -0.394764, but must be positive! (in '/tmp/RtmpjPjDJw/model-122683fea6.stan', line 18, column 4 to column 29)
Chain 3 Exception: normal_lpdf: Scale parameter is -0.394764, but must be positive! (in '/tmp/RtmpjPjDJw/model-122683fea6.stan', line 18, column 4 to column 29)
Chain 3 Rejecting initial value:
Chain 3 Error evaluating the log probability at the initial value.
Chain 3 Exception: normal_lpdf: Scale parameter is -1.13481, but must be positive! (in '/tmp/RtmpjPjDJw/model-122683fea6.stan', line 18, column 4 to column 29)
Chain 3 Exception: normal_lpdf: Scale parameter is -1.13481, but must be positive! (in '/tmp/RtmpjPjDJw/model-122683fea6.stan', line 18, column 4 to column 29)
Chain 3 Rejecting initial value:
Chain 3 Error evaluating the log probability at the initial value.
Chain 3 Exception: normal_lpdf: Scale parameter is -0.175318, but must be positive! (in '/tmp/RtmpjPjDJw/model-122683fea6.stan', line 18, column 4 to column 29)
Chain 3 Exception: normal_lpdf: Scale parameter is -0.175318, but must be positive! (in '/tmp/RtmpjPjDJw/model-122683fea6.stan', line 18, column 4 to column 29)
Chain 3 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 3 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 3 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 3 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 3 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 3 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 3 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 3 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 3 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 3 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 3 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 3 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 3 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 3 Exception: normal_lpdf: Scale parameter is -0.333645, but must be positive! (in '/tmp/RtmpjPjDJw/model-122683fea6.stan', line 18, column 4 to column 29)
Chain 3 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 3 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 3
Chain 3 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 3 Exception: normal_lpdf: Scale parameter is -0.819312, but must be positive! (in '/tmp/RtmpjPjDJw/model-122683fea6.stan', line 18, column 4 to column 29)
Chain 3 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 3 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 3
Chain 4 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 4 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 4 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 4 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 4 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 4 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 4 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 4 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 4 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 4 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 4 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 4 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 4 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 4 Exception: normal_lpdf: Scale parameter is -1.57384, but must be positive! (in '/tmp/RtmpjPjDJw/model-122683fea6.stan', line 18, column 4 to column 29)
Chain 4 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 4 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 4
Chain 4 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 4 Exception: normal_lpdf: Scale parameter is -0.695607, but must be positive! (in '/tmp/RtmpjPjDJw/model-122683fea6.stan', line 18, column 4 to column 29)
Chain 4 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 4 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 4
Chain 1 finished in 0.1 seconds.
Chain 2 finished in 0.0 seconds.
Chain 3 finished in 0.0 seconds.
Chain 4 finished in 0.0 seconds.
All 4 chains finished successfully.
Mean chain execution time: 0.0 seconds.
Total execution time: 0.4 seconds.
Raw data (following plot):
mean sd 5.5% 94.5% n_eff Rhat4
b 1.0796106 0.4921676 0.2869450 1.8460359 1080.7259 1.0021330
a 0.2105812 0.2997821 -0.2759974 0.6750308 1180.1113 0.9997719
sigma 0.9642299 0.2458819 0.6491778 1.3962750 877.1299 1.0038247
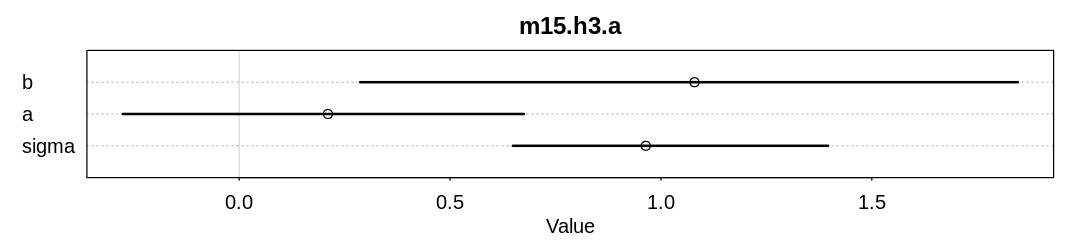
Fitting a model to all the data, with the imputation model suggested in the question:
q.15.h3.b <- function(d) {
dat <- list(
Y = d$y,
X = d$x
)
m15.h3.b <- ulam(
alist(
Y ~ dnorm(mu, sigma),
mu <- a + b*X,
X ~ dnorm(0, 1),
c(a, b) ~ dnorm(0, 1),
sigma ~ dexp(1)
), cmdstan = TRUE, data=dat, cores=4, chains=4, log_lik=TRUE
)
display_precis(m15.h3.b, "m15.h3.b", 4.0)
return(m15.h3.b)
}
m15.h3.b <- q.15.h3.b(d.15.h3)
Found 1 NA values in X and attempting imputation.
Warning in '/tmp/RtmpjPjDJw/model-12446ee951.stan', line 4, column 26: Declaration
of arrays by placing brackets after a type is deprecated and will be
removed in Stan 2.32.0. Instead use the array keyword before the type.
This can be changed automatically using the auto-format flag to stanc
Warning in '/tmp/RtmpjPjDJw/model-12446ee951.stan', line 17, column 4: Declaration
of arrays by placing brackets after a variable name is deprecated and
will be removed in Stan 2.32.0. Instead use the array keyword before the
type. This can be changed automatically using the auto-format flag to
stanc
Running MCMC with 4 parallel chains, with 1 thread(s) per chain...
Chain 1 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 1 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 1 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 1 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 1 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 1 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 1 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 1 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 1 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 1 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 1 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 1 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 2 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 2 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 2 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 2 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 2 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 2 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 2 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 2 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 2 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 2 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 2 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 2 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 3 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 3 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 3 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 3 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 3 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 3 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 3 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 3 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 3 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 3 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 3 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 3 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 4 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 4 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 4 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 4 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 4 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 4 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 4 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 4 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 4 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 4 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 4 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 4 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 1 finished in 0.1 seconds.
Chain 2 finished in 0.1 seconds.
Chain 3 finished in 0.1 seconds.
Chain 4 finished in 0.1 seconds.
All 4 chains finished successfully.
Mean chain execution time: 0.1 seconds.
Total execution time: 0.2 seconds.
Raw data (following plot):
mean sd 5.5% 94.5% n_eff Rhat4
b 0.02013376 1.0382012 -1.684889 1.686190 2102.224 0.9996513
a 0.29524993 0.9979384 -1.314590 1.880945 2317.508 0.9989613
sigma 18.87964020 2.2106872 15.649452 22.568184 2517.860 1.0004584
X_impute[1] -0.01639253 1.0044745 -1.625428 1.616405 2165.268 0.9994991
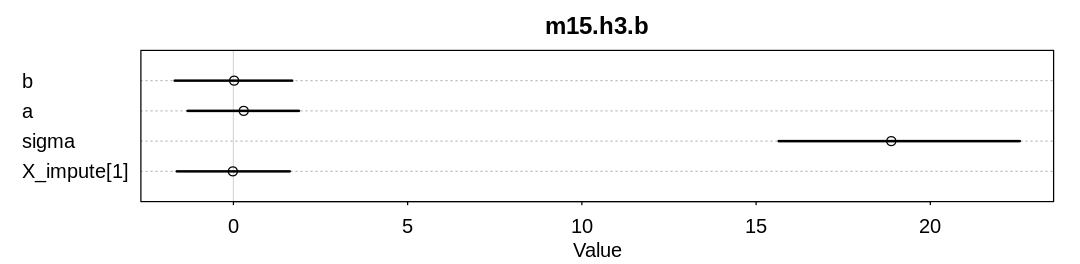
Notice the model has imputed a value near zero for the single missing value. This point is extremely
influential because of the large value of y associated with it (the 100) so it dominates the
inference results. A value of zero associated with 100 implies almost no relationship between x
and y so the model infers that sigma is large instead.
15H4. Using data(Primates301), consider the relationship between brain volume (brain) and
body mass (body). These variables are presented as single values for each species. However, there
is always a range of sizes in a species, and some of these measurements are taken from very small
samples. So these values are measured with some unknown error.
We don’t have the raw measurements to work with - that would be best. But we can imagine what might happen if we had them. Suppose error is proportional to the measurement. This makes sense, because larger animals have larger variation. As a consequence, the uncertainty is not uniform across the values and this could mean trouble.
Let’s make up some standard errors for these measurements, to see what might happen. Load the data and scale the the measurements so the maximum is 1 in both cases:
R code 15.33
library(rethinking)
data(Primates301)
d <- Primates301
cc <- complete.cases( d$brain , d$body )
B <- d$brain[cc]
M <- d$body[cc]
B <- B / max(B)
M <- M / max(M)
Now I’ll make up some standard errors for \(B\) and \(M\), assuming error is 10% of the measurement.
R code 15.34
Bse <- B*0.1
Mse <- M*0.1
Let’s model these variables with this relationship:
This says that brain volume is a log-normal variable, and the mean on the log scale is given by \(\mu\). What this model implies is that the expected value of B is:
So this is a standard allometric scaling relationship - incredibly common in biology.
Ignoring measurement error, the corresponding ulam model is:
R code 15.35
dat_list <- list( B = B , M = M )
m15H4 <- ulam(
alist(
B ~ dlnorm( mu , sigma ),
mu <- a + b*log(M),
a ~ normal(0,1),
b ~ normal(0,1),
sigma ~ exponential(1)
) , cmdstan = TRUE, data=dat_list )
Your job is to add the measurement errors to this model. Use the divorce/marriage example in the
chapter as a guide. It might help to initialize the unobserved true values of \(B\) and \(M\) using the
observed values, by adding a list like this to ulam:
R code 15.36
start=list( M_true=dat_list$M , B_true=dat_list$B )
Compare the inference of the measurement error model to those of m1.1 above. Has anything changed?
Why or why not?
ERROR:
Compare the inference of the measurement error model to those of
m1.1above.
The author meant m15H4.
Solution. The math is a bit fuzzy for the allometric relationship. The mean of a LogNormal random variable is:
The author seems to be ignoring the \(\sigma^2/2\) term. If you accept that ‘simplification’ you can get the second equation he presents with:
load.d.15.h4 <- function() {
data(Primates301)
d <- Primates301
cc <- complete.cases( d$brain , d$body )
B <- d$brain[cc]
M <- d$body[cc]
B <- B / max(B)
M <- M / max(M)
Bse <- B*0.1
Mse <- M*0.1
d.15.h4 <- data.frame(B=B, M=M, Bse=Bse, Mse=Mse)
return(d.15.h4)
}
d.15.h4 <- load.d.15.h4()
q.15.h4.a <- function(d) {
dat_list <- list(B = d$B, M = d$M)
m15H4 <- ulam(
alist(
B ~ dlnorm(mu, sigma),
mu <- a + b * log(M),
a ~ normal(0, 1),
b ~ normal(0, 1),
sigma ~ exponential(1)
), cmdstan = TRUE, data=dat_list, chains=4, cores=4
)
display_precis(m15H4, "m15H4", 4.0)
return(m15H4)
}
m15H4 <- q.15.h4.a(d.15.h4)
q.15.h4.b <- function(d) {
dat_list <- list(
B_obs = d$B,
M_obs = d$M,
B_sd = d$Bse,
M_sd = d$Mse,
N_obs = nrow(d)
)
m.15.h4.b <- ulam(
alist(
B_obs ~ dnorm(B_true, B_sd),
vector[N_obs]:B_true ~ dlnorm(mu, sigma),
mu <- a + b * log(M_true[i]),
M_obs ~ dnorm(M_true, M_sd),
vector[N_obs]:M_true ~ dexp(1),
a ~ normal(0, 1),
b ~ normal(0, 1),
sigma ~ exponential(1)
), cmdstan = TRUE, data=dat_list, chains=4, cores=4, log_lik=TRUE
)
display_precis(m.15.h4.b, "m.15.h4.b", 0.2)
return(m.15.h4.b)
}
m.15.h4.b <- q.15.h4.b(d.15.h4)
# iplot_coeftab <- function(..., pars, ar, main="coeftab") {
# iplot(function() {
# plot(coeftab(...), pars=pars, main=main)
# }, ar=ar)
# }
# iplot_coeftab(m15H4, m.15.h4.b, pars=c("a", "b", "sigma"), ar=2.0)
iplot(function() {
plot(coeftab(m15H4, m.15.h4.b), pars=c("a", "b", "sigma"), main="coeftab")
}, ar=2)
Running MCMC with 4 parallel chains, with 1 thread(s) per chain...
Chain 1 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 1 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 1 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 1 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 1 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 1 Exception: lognormal_lpdf: Scale parameter is 0, but must be positive finite! (in '/tmp/RtmpjPjDJw/model-121c107d98.stan', line 18, column 4 to column 32)
Chain 1 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 1 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 1
Chain 2 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 2 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 2 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 2 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 2 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 2 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 2 Exception: lognormal_lpdf: Scale parameter is 0, but must be positive finite! (in '/tmp/RtmpjPjDJw/model-121c107d98.stan', line 18, column 4 to column 32)
Chain 2 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 2 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 2
Chain 3 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 3 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 3 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 3 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 3 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 4 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 4 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 4 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 4 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 4 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 4 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 4 Exception: lognormal_lpdf: Scale parameter is inf, but must be positive finite! (in '/tmp/RtmpjPjDJw/model-121c107d98.stan', line 18, column 4 to column 32)
Chain 4 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 4 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 4
Chain 1 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 1 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 1 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 1 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 1 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 1 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 1 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 1 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 2 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 2 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 2 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 2 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 2 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 2 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 2 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 3 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 3 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 3 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 3 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 3 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 3 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 4 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 4 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 4 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 4 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 4 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 4 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 1 finished in 0.3 seconds.
Chain 2 finished in 0.3 seconds.
Chain 3 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 4 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 3 finished in 0.3 seconds.
Chain 4 finished in 0.3 seconds.
All 4 chains finished successfully.
Mean chain execution time: 0.3 seconds.
Total execution time: 0.5 seconds.
Raw data (following plot):
mean sd 5.5% 94.5% n_eff Rhat4
a 0.4264338 0.05988520 0.3312353 0.5224070 700.3517 1.003542
b 0.7830678 0.01441620 0.7598551 0.8058107 711.1302 1.003888
sigma 0.2936612 0.01548044 0.2704216 0.3199446 1012.5430 1.000912
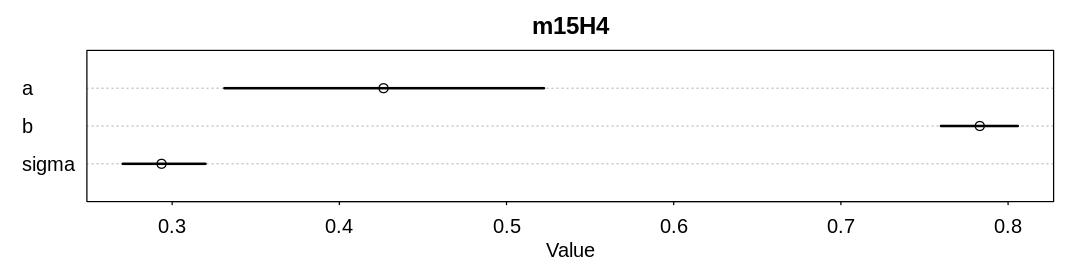
Running MCMC with 4 parallel chains, with 1 thread(s) per chain...
Chain 1 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 1 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 1 Exception: lognormal_lpdf: Location parameter[1] is -inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 1 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 1 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 1
Chain 1 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 1 Exception: lognormal_lpdf: Location parameter[1] is -inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 1 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 1 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 1
Chain 1 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 1 Exception: lognormal_lpdf: Location parameter[2] is -inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 1 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 1 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 1
Chain 1 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 1 Exception: lognormal_lpdf: Location parameter[2] is inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 1 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 1 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 1
Chain 1 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 1 Exception: lognormal_lpdf: Location parameter[2] is -inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 1 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 1 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 1
Chain 1 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 1 Exception: lognormal_lpdf: Location parameter[2] is -inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 1 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 1 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 1
Chain 1 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 1 Exception: lognormal_lpdf: Location parameter[9] is -inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 1 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 1 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 1
Chain 1 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 1 Exception: lognormal_lpdf: Location parameter[9] is -inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 1 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 1 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 1
Chain 1 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 1 Exception: lognormal_lpdf: Location parameter[9] is -inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 1 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 1 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 1
Chain 1 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 1 Exception: lognormal_lpdf: Location parameter[25] is -inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 1 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 1 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 1
Chain 1 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 1 Exception: lognormal_lpdf: Location parameter[74] is -inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 1 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 1 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 1
Chain 1 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 1 Exception: lognormal_lpdf: Location parameter[79] is -inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 1 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 1 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 1
Chain 1 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 1 Exception: normal_lpdf: Location parameter[81] is inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 21, column 4 to column 36)
Chain 1 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 1 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 1
Chain 1 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 1 Exception: lognormal_lpdf: Location parameter[1] is -inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 1 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 1 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 1
Chain 1 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 1 Exception: lognormal_lpdf: Location parameter[2] is -inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 1 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 1 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 1
Chain 1 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 1 Exception: lognormal_lpdf: Location parameter[25] is -inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 1 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 1 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 1
Chain 2 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 2 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 2 Exception: lognormal_lpdf: Location parameter[1] is inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 2 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 2 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 2
Chain 2 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 2 Exception: lognormal_lpdf: Location parameter[1] is inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 2 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 2 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 2
Chain 2 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 2 Exception: lognormal_lpdf: Location parameter[1] is inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 2 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 2 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 2
Chain 2 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 2 Exception: lognormal_lpdf: Location parameter[1] is inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 2 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 2 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 2
Chain 2 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 2 Exception: lognormal_lpdf: Location parameter[1] is inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 2 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 2 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 2
Chain 2 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 2 Exception: lognormal_lpdf: Location parameter[1] is inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 2 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 2 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 2
Chain 2 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 2 Exception: lognormal_lpdf: Location parameter[10] is inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 2 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 2 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 2
Chain 2 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 2 Exception: lognormal_lpdf: Location parameter[10] is inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 2 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 2 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 2
Chain 2 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 2 Exception: lognormal_lpdf: Location parameter[10] is inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 2 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 2 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 2
Chain 2 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 2 Exception: lognormal_lpdf: Location parameter[10] is inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 2 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 2 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 2
Chain 2 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 2 Exception: lognormal_lpdf: Location parameter[27] is inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 2 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 2 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 2
Chain 2 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 2 Exception: lognormal_lpdf: Location parameter[27] is inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 2 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 2 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 2
Chain 2 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 2 Exception: normal_lpdf: Location parameter[81] is inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 21, column 4 to column 36)
Chain 2 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 2 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 2
Chain 2 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 2 Exception: lognormal_lpdf: Location parameter[1] is inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 2 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 2 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 2
Chain 2 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 2 Exception: lognormal_lpdf: Location parameter[1] is inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 2 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 2 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 2
Chain 2 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 2 Exception: lognormal_lpdf: Location parameter[10] is inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 2 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 2 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 2
Chain 3 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 3 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 3 Exception: lognormal_lpdf: Location parameter[1] is inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 3 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 3 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 3
Chain 3 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 3 Exception: lognormal_lpdf: Location parameter[1] is -inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 3 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 3 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 3
Chain 3 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 3 Exception: lognormal_lpdf: Location parameter[1] is -inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 3 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 3 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 3
Chain 3 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 3 Exception: lognormal_lpdf: Location parameter[1] is -inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 3 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 3 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 3
Chain 3 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 3 Exception: lognormal_lpdf: Location parameter[1] is -inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 3 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 3 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 3
Chain 3 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 3 Exception: lognormal_lpdf: Location parameter[5] is -inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 3 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 3 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 3
Chain 3 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 3 Exception: lognormal_lpdf: Location parameter[8] is -inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 3 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 3 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 3
Chain 3 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 3 Exception: lognormal_lpdf: Location parameter[11] is -inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 3 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 3 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 3
Chain 3 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 3 Exception: lognormal_lpdf: Location parameter[27] is -inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 3 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 3 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 3
Chain 3 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 3 Exception: lognormal_lpdf: Location parameter[79] is -inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 3 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 3 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 3
Chain 3 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 3 Exception: lognormal_lpdf: Location parameter[79] is -inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 3 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 3 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 3
Chain 3 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 3 Exception: normal_lpdf: Location parameter[81] is inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 21, column 4 to column 36)
Chain 3 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 3 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 3
Chain 3 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 3 Exception: lognormal_lpdf: Location parameter[1] is -inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 3 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 3 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 3
Chain 3 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 3 Exception: lognormal_lpdf: Location parameter[5] is -inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 3 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 3 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 3
Chain 4 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 4 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 4 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 4 Exception: lognormal_lpdf: Location parameter[1] is -inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 4 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 4 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 4
Chain 4 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 4 Exception: lognormal_lpdf: Location parameter[1] is -inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 4 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 4 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 4
Chain 4 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 4 Exception: lognormal_lpdf: Location parameter[1] is -inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 4 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 4 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 4
Chain 4 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 4 Exception: lognormal_lpdf: Location parameter[3] is inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 4 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 4 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 4
Chain 4 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 4 Exception: lognormal_lpdf: Location parameter[3] is inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 4 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 4 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 4
Chain 4 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 4 Exception: lognormal_lpdf: Location parameter[4] is inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 4 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 4 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 4
Chain 4 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 4 Exception: lognormal_lpdf: Location parameter[4] is inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 4 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 4 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 4
Chain 4 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 4 Exception: lognormal_lpdf: Location parameter[9] is inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 4 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 4 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 4
Chain 4 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 4 Exception: lognormal_lpdf: Location parameter[9] is inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 4 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 4 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 4
Chain 4 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 4 Exception: lognormal_lpdf: Location parameter[13] is inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 4 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 4 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 4
Chain 4 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 4 Exception: lognormal_lpdf: Location parameter[13] is inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 4 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 4 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 4
Chain 4 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 4 Exception: lognormal_lpdf: Location parameter[54] is inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 4 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 4 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 4
Chain 4 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 4 Exception: normal_lpdf: Location parameter[81] is inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 21, column 4 to column 36)
Chain 4 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 4 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 4
Chain 4 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 4 Exception: lognormal_lpdf: Location parameter[1] is -inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 4 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 4 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 4
Chain 4 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 4 Exception: lognormal_lpdf: Location parameter[3] is inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 4 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 4 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 4
Chain 4 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 4 Exception: lognormal_lpdf: Location parameter[13] is inf, but must be finite! (in '/tmp/RtmpjPjDJw/model-127854c061.stan', line 25, column 4 to column 37)
Chain 4 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 4 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 4
Chain 1 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 1 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 1 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 1 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 2 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 2 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 3 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 3 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 4 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 1 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 3 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 4 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 1 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 2 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 1 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 2 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 3 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 4 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 1 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 3 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 3 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 4 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 4 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 2 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 2 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 1 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 3 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 4 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 2 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 1 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 2 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 3 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 4 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 1 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 3 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 4 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 1 finished in 2.1 seconds.
Chain 2 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 3 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 4 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 2 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 2 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 3 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 4 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 2 finished in 2.5 seconds.
Chain 3 finished in 2.4 seconds.
Chain 4 finished in 2.4 seconds.
All 4 chains finished successfully.
Mean chain execution time: 2.3 seconds.
Total execution time: 2.5 seconds.
Raw data (following plot):
mean sd 5.5% 94.5% n_eff
B_true[1] 0.116501049 0.0108402991 0.098894523 0.133888925 4151.292
B_true[2] 0.110231006 0.0101802896 0.094368209 0.126886765 4166.197
B_true[3] 0.108207786 0.0102709931 0.092084153 0.124479100 5120.396
B_true[4] 0.106226831 0.0095893161 0.090907526 0.121450265 4162.346
B_true[5] 0.104855479 0.0098984460 0.089063354 0.121058310 4432.129
B_true[6] 0.110209395 0.0094850710 0.094820756 0.125500275 4889.156
B_true[7] 0.122010135 0.0120237453 0.102857645 0.141140965 4963.691
B_true[8] 0.114012183 0.0100984946 0.097958978 0.130092905 5001.726
B_true[9] 0.041210848 0.0039537034 0.034655030 0.047464372 4923.059
B_true[10] 0.031967147 0.0030520460 0.027005560 0.036873166 4120.390
B_true[11] 0.033990621 0.0032671643 0.028946007 0.039204179 5147.285
B_true[12] 0.011615569 0.0011530438 0.009795893 0.013472539 3759.581
B_true[13] 0.013879664 0.0013195969 0.011810195 0.016042645 5508.006
B_true[14] 0.226184862 0.0229068566 0.189930505 0.262839200 5025.154
B_true[15] 0.224777534 0.0220302396 0.189755270 0.259387860 4569.248
B_true[16] 0.205150567 0.0197638475 0.174193665 0.236584235 4188.408
B_true[17] 0.205278207 0.0203633079 0.171790135 0.238753335 5450.778
B_true[18] 0.021432650 0.0018572777 0.018437834 0.024353876 4434.788
B_true[19] 0.017127153 0.0015081086 0.014662933 0.019543994 4186.990
B_true[20] 0.211951499 0.0227170391 0.174935600 0.247565000 5339.681
B_true[21] 0.140854892 0.0151150537 0.116608590 0.164992495 4918.582
B_true[22] 0.128264715 0.0137268858 0.106459525 0.150669660 4512.236
B_true[23] 0.022477453 0.0021018929 0.019144861 0.025896720 4993.448
B_true[24] 0.015836114 0.0014732390 0.013535502 0.018186942 5526.985
B_true[25] 0.014493791 0.0014178371 0.012279597 0.016777236 6602.060
B_true[26] 0.014668672 0.0013989743 0.012521752 0.016804271 4540.035
B_true[27] 0.008086627 0.0007714216 0.006856515 0.009321702 6237.143
B_true[28] 0.121937390 0.0125656239 0.102376120 0.141602715 4603.340
B_true[29] 0.124567333 0.0134158195 0.103564470 0.146810950 4432.053
B_true[30] 0.134406476 0.0142944320 0.111640525 0.157261400 3971.116
⋮ ⋮ ⋮ ⋮ ⋮ ⋮
M_true[156] 0.0218681341 2.198526e-03 0.0184078130 0.025338225 4455.942
M_true[157] 0.0740924054 7.433089e-03 0.0621375165 0.086132004 4547.262
M_true[158] 0.1119229651 1.180245e-02 0.0931663405 0.130695530 5311.109
M_true[159] 0.0030852139 2.975357e-04 0.0026144887 0.003569972 4734.040
M_true[160] 0.0039918902 3.786749e-04 0.0033638231 0.004593379 4221.715
M_true[161] 0.0040281731 4.056766e-04 0.0033881246 0.004692708 5922.510
M_true[162] 0.0043007993 4.155236e-04 0.0036304378 0.004958710 3708.773
M_true[163] 0.0045064638 4.245070e-04 0.0038333670 0.005177733 4897.942
M_true[164] 0.0029434441 2.700648e-04 0.0025215720 0.003363746 5842.937
M_true[165] 0.0033507184 3.271177e-04 0.0028392695 0.003860236 4454.267
M_true[166] 0.0063850683 5.773970e-04 0.0054606781 0.007314350 4608.478
M_true[167] 0.0064372238 5.972085e-04 0.0054688097 0.007427873 5125.998
M_true[168] 0.1110327053 1.096326e-02 0.0936650190 0.128780385 5002.494
M_true[169] 0.0878963590 8.510442e-03 0.0745290905 0.101560850 5301.929
M_true[170] 0.0009667255 9.704217e-05 0.0008089893 0.001120210 4625.569
M_true[171] 0.0008680478 8.382024e-05 0.0007257090 0.001002774 3178.937
M_true[172] 0.0009747715 9.358022e-05 0.0008231044 0.001120076 4790.759
M_true[173] 0.1173411332 1.179082e-02 0.0990117515 0.136820390 4913.305
M_true[174] 0.0481464885 5.088016e-03 0.0399478245 0.056162464 4459.045
M_true[175] 0.0764081500 7.633404e-03 0.0643051320 0.088423083 5720.210
M_true[176] 0.0866757924 8.740503e-03 0.0729003225 0.100575090 5683.205
M_true[177] 0.0530617235 5.169180e-03 0.0448247865 0.061317172 3356.499
M_true[178] 0.0569924843 5.634844e-03 0.0477599575 0.065910313 4385.082
M_true[179] 0.0898655397 8.890999e-03 0.0755746165 0.104114510 5428.557
M_true[180] 0.0474043347 4.704109e-03 0.0398315630 0.054710805 4639.041
M_true[181] 0.0258553452 2.651107e-03 0.0215936340 0.030070616 4128.754
M_true[182] 0.0265882314 2.706218e-03 0.0222069710 0.030818392 5536.600
a 0.4191816405 5.932353e-02 0.3262413600 0.513002280 2767.671
b 0.7838324045 1.417906e-02 0.7607243800 0.806090665 2770.061
sigma 0.2627569540 1.680301e-02 0.2372583950 0.290605660 2309.916
Rhat4
B_true[1] 0.9985442
B_true[2] 0.9997402
B_true[3] 0.9983380
B_true[4] 0.9990252
B_true[5] 0.9981828
B_true[6] 0.9987771
B_true[7] 0.9984331
B_true[8] 0.9983743
B_true[9] 0.9992680
B_true[10] 0.9983893
B_true[11] 0.9986528
B_true[12] 1.0000221
B_true[13] 0.9983626
B_true[14] 0.9982209
B_true[15] 0.9983817
B_true[16] 0.9990960
B_true[17] 0.9988796
B_true[18] 0.9992431
B_true[19] 0.9995469
B_true[20] 0.9984028
B_true[21] 0.9990997
B_true[22] 0.9989766
B_true[23] 0.9993852
B_true[24] 0.9986114
B_true[25] 0.9985109
B_true[26] 0.9993952
B_true[27] 0.9986008
B_true[28] 0.9988681
B_true[29] 0.9993563
B_true[30] 0.9993881
⋮ ⋮
M_true[156] 0.9992022
M_true[157] 0.9989419
M_true[158] 0.9991944
M_true[159] 0.9983307
M_true[160] 0.9986911
M_true[161] 0.9986974
M_true[162] 0.9994798
M_true[163] 0.9998614
M_true[164] 0.9984022
M_true[165] 0.9991358
M_true[166] 0.9987511
M_true[167] 0.9982739
M_true[168] 0.9986564
M_true[169] 0.9987730
M_true[170] 0.9986466
M_true[171] 1.0001176
M_true[172] 0.9992279
M_true[173] 0.9987013
M_true[174] 0.9988869
M_true[175] 0.9985858
M_true[176] 0.9982319
M_true[177] 0.9990161
M_true[178] 0.9993060
M_true[179] 0.9996466
M_true[180] 0.9993738
M_true[181] 0.9995555
M_true[182] 0.9989652
a 0.9989791
b 0.9991454
sigma 1.0007652
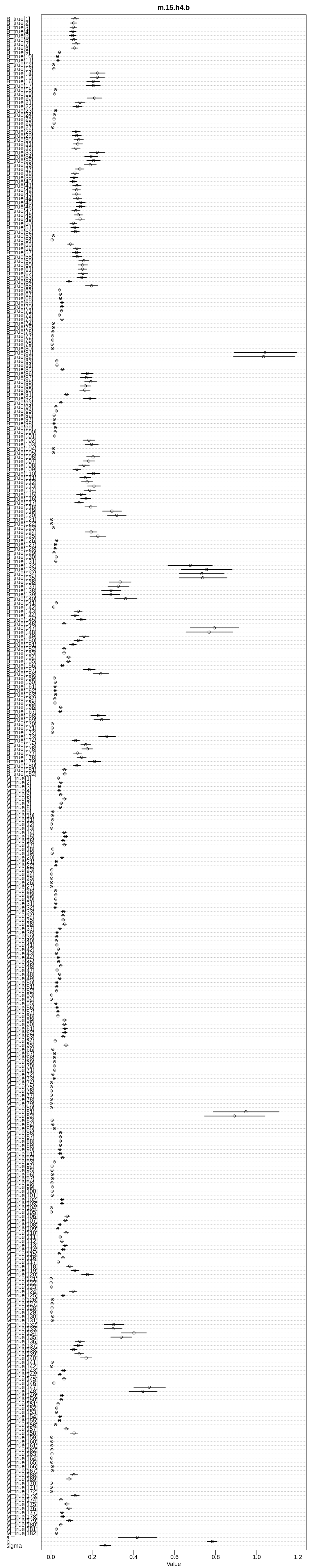
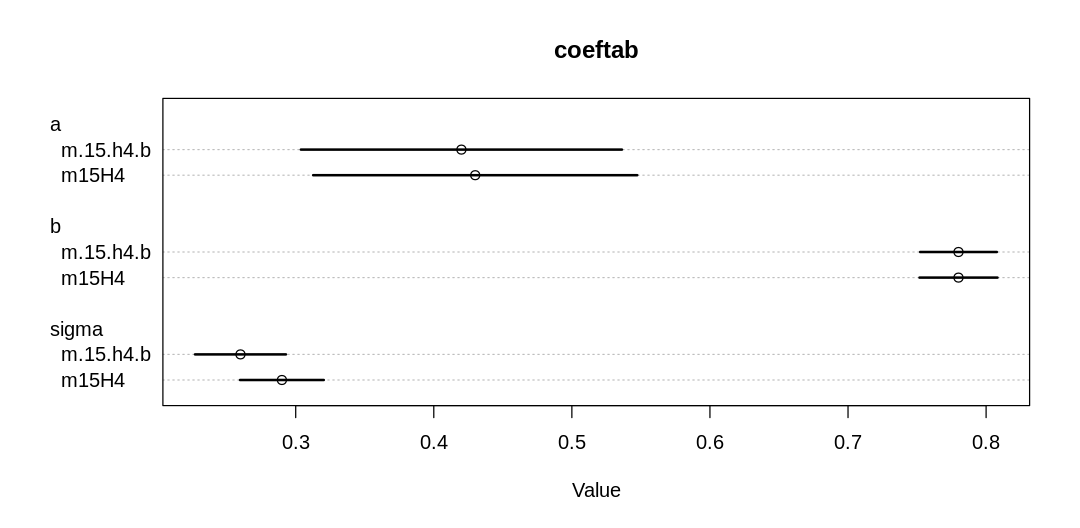
Little has changed with the update. It looks like sigma is slightly smaller, perhaps because more
uncertainty in the data has naturally led to more confidence in the model.
15H5. Now consider missing values - this data set is lousy with them. You can ignore measurement error in this problem. Let’s get a quick idea of the missing values by counting them in each variable: R code
R code 15.37
library(rethinking)
data(Primates301)
d <- Primates301
colSums( is.na(d) )
We’ll continue to focus on just brain and body, to stave off insanity. Consider only those species with measured body masses:
R code 15.38
cc <- complete.cases( d$body )
M <- d$body[cc]
M <- M / max(M)
B <- d$brain[cc]
B <- B / max( B , na.rm=TRUE )
You should end up with 238 species and 56 missing brain values among them.
First, consider whether there is a pattern to the missing values. Does it look like missing values are associated with particular values of body mass? Draw a DAG that represents how missingness works in this case. Which type (MCAR, MAR, MNAR) is this?
Second, impute missing values for brain size. It might help to initialize the 56 imputed variables to a valid value:
R code 15.39
start=list( B_impute=rep(0.5,56) )
This just helps the chain get started.
Compare the inferences to an analysis that drops all the missing values. Has anything changed? Why or why not? Hint: Consider the density of data in the ranges where there are missing values. You might want to plot the imputed brain sizes together with the observed values.
ERROR:
by counting them in each variable: R code
load.d15h5 <- function() {
data(Primates301)
d <- Primates301
display_markdown("**Answer.** The `colSums` output:")
display(colSums(is.na(d)), mimetypes="text/plain")
cc <- complete.cases( d$body )
M <- d$body[cc]
M <- M / max(M)
B <- d$brain[cc]
B <- B / max( B , na.rm=TRUE )
d15h5 <- data.frame(B = B, M = M)
display_markdown("<br/> To confirm there are 238 species, preview the data.frame:")
display(d15h5)
display_markdown("<br/> To confirm there are 56 missing brain values, check the `summary`:")
display(summary(d15h5))
return(d15h5)
}
e15h5a <- r"(
The negative intercept `a` merely indicates that most values are 0 rather than 1 in D (not missing).
More interesting is `bM`, which is reliably negative. A negative `bM` indicates that larger values
of `M` are associated with being missing less. That is, brain sizes are missing more often for
species with small masses.
Clearly this is not MCAR. See section **15.2.2** for potential DAGs. If you believe that brain
volumes are only missing because researchers prefer to study species with a larger mass, then this
may be a salvageable MAR situation. If you believe brain volumes are missing because researchers
prefer to study species with larger brains, this is MNAR.
)"
q15h5a <- function(d) {
dat <- list(
D = as.integer(is.na(d$B)),
M = as.numeric(standardize(d$M))
)
display_markdown("<br/> Let's check if we can predict `D` (deleted values) from `M`:")
m15h5a <- ulam(
alist(
D ~ dbinom(1, p),
logit(p) <- a + bM*M,
c(a, bM) ~ dnorm(0, 1)
), cmdstan = TRUE, data=dat, cores=4, chains=4
)
display_precis(m15h5a, "m15h5a", ar=4.0)
display_markdown(e15h5a)
}
q15h5b <- function(d) {
dat <- list(
B = d$B,
M = d$M
)
# browser()
m15h5b <- ulam(
alist(
B ~ dlnorm(mu, sigma),
mu <- a + b * log(M),
a ~ normal(0, 1),
b ~ normal(0, 1),
sigma ~ exponential(1)
), cmdstan=TRUE, data=dat, cores=4, chains=4, start=list( B_impute=rep(0.5,56) )
)
display_precis(m15h5b, "m15h5b", ar=0.7)
return(m15h5b)
}
q15h5cc <- function(d) {
d_cc <- d[complete.cases(d$B),]
dat <- list(B = d_cc$B, M = d_cc$M)
m15H4cc <- ulam(
alist(
B ~ dlnorm(mu, sigma),
mu <- a + b * log(M),
a ~ normal(0, 1),
b ~ normal(0, 1),
sigma ~ exponential(1)
), cmdstan=TRUE, data=dat, chains=4, cores=4
)
display_precis(m15H4cc, "m15H4cc", 4.0)
return(m15H4cc)
}
e15h5 <- r"(
There doesn't seem to be any change in the inferences for `a`, `b`, and `sigma`. When you impute
output values, the model isn't really doing anything but posterior predictive inference. We haven't
supplied any new assumptions about what the distribution of `B` should look like, other than what we
know from `M` (which we only learned from complete cases).
)"
q15h5 <- function() {
d15h5 <- load.d15h5()
q15h5a(d15h5)
m15h5b <- q15h5b(d15h5)
m15H4cc <- q15h5cc(d15h5)
iplot(function() {
plot(coeftab(m15H4cc, m15h5b), pars=c("a", "b", "sigma"), main="coeftab")
}, ar=2)
display_markdown(e15h5)
}
q15h5()
Answer. The colSums output:
name genus species subspecies
0 0 0 267
spp_id genus_id social_learning research_effort
0 0 98 115
brain body group_size gestation
117 63 114 161
weaning longevity sex_maturity maternal_investment
185 181 194 197
To confirm there are 238 species, preview the data.frame:
| B | M |
|---|---|
| <dbl> | <dbl> |
| 0.11810206 | 0.0358076923 |
| NA | 0.0006006923 |
| 0.10755796 | 0.0491923077 |
| 0.10713050 | 0.0414076923 |
| 0.10523745 | 0.0398076923 |
| 0.10153276 | 0.0480769231 |
| 0.10407719 | 0.0685769231 |
| 0.12025973 | 0.0508541538 |
| 0.11240255 | 0.0457692308 |
| 0.04207462 | 0.0092692308 |
| 0.03317931 | 0.0056461538 |
| NA | 0.0060848462 |
| NA | 0.0073692308 |
| 0.03429886 | 0.0076076923 |
| NA | 0.0054076923 |
| 0.01196898 | 0.0016153846 |
| 0.01408594 | 0.0023769231 |
| 0.23819895 | 0.0628230769 |
| 0.23254015 | 0.0694230769 |
| 0.21391496 | 0.0579615385 |
| 0.21139088 | 0.0636923077 |
| 0.02007043 | 0.0092846154 |
| 0.01618255 | 0.0061615385 |
| NA | 0.0810562308 |
| 0.22529363 | 0.0517538462 |
| 0.15470108 | 0.0243461538 |
| 0.13998412 | 0.0225769231 |
| NA | 0.0069051538 |
| NA | 0.0082123846 |
| NA | 0.0073702308 |
| ⋮ | ⋮ |
| 0.016162192 | 0.0030846154 |
| 0.020640381 | 0.0039769231 |
| NA | 0.0031377692 |
| 0.019744743 | 0.0040384615 |
| 0.019907586 | 0.0043307692 |
| 0.022574145 | 0.0044923077 |
| 0.019296924 | 0.0028846154 |
| 0.019866876 | 0.0033153846 |
| NA | 0.0029619231 |
| NA | 0.0061496154 |
| 0.051031001 | 0.0060692308 |
| 0.049137949 | 0.0061461538 |
| NA | 0.0068190000 |
| 0.225802512 | 0.1134000000 |
| 0.251389256 | 0.0868846154 |
| 0.006432308 | 0.0009692308 |
| 0.006106622 | 0.0008692308 |
| 0.006839416 | 0.0009692308 |
| 0.271398620 | 0.1180769231 |
| NA | 0.0747661538 |
| 0.117776376 | 0.0491846154 |
| NA | 0.0626148462 |
| 0.165489446 | 0.0780769231 |
| 0.172206729 | 0.0892307692 |
| 0.126447778 | 0.0542769231 |
| 0.148268773 | 0.0575000000 |
| 0.210963421 | 0.0907230769 |
| 0.124758280 | 0.0479769231 |
| 0.063264600 | 0.0266923077 |
| 0.065381562 | 0.0275000000 |
To confirm there are 56 missing brain values, check the summary:
B M
Min. :0.00332 Min. :0.0002402
1st Qu.:0.02440 1st Qu.:0.0056880
Median :0.12075 Median :0.0273346
Mean :0.14075 Mean :0.0522706
3rd Qu.:0.17789 3rd Qu.:0.0574231
Max. :1.00000 Max. :1.0000000
NA's :56
Let’s check if we can predict D (deleted values) from M:
Warning in '/tmp/RtmpjPjDJw/model-122d3b96cb.stan', line 2, column 4: Declaration
of arrays by placing brackets after a variable name is deprecated and
will be removed in Stan 2.32.0. Instead use the array keyword before the
type. This can be changed automatically using the auto-format flag to
stanc
Running MCMC with 4 parallel chains, with 1 thread(s) per chain...
Chain 1 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 1 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 1 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 1 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 1 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 1 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 1 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 1 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 1 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 1 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 2 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 2 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 2 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 2 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 2 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 2 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 2 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 2 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 2 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 2 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 2 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 2 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 3 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 3 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 3 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 3 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 3 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 3 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 3 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 3 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 3 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 3 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 3 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 3 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 4 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 4 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 4 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 4 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 4 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 4 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 4 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 4 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 1 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 1 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 1 finished in 0.2 seconds.
Chain 2 finished in 0.2 seconds.
Chain 3 finished in 0.2 seconds.
Chain 4 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 4 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 4 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 4 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 4 finished in 0.2 seconds.
All 4 chains finished successfully.
Mean chain execution time: 0.2 seconds.
Total execution time: 0.4 seconds.
Raw data (following plot):
mean sd 5.5% 94.5% n_eff Rhat4
bM -0.5072257 0.3028944 -1.048434 -0.06574995 1014.347 0.9996125
a -1.2086073 0.1610078 -1.474740 -0.95873821 1127.009 0.9996780
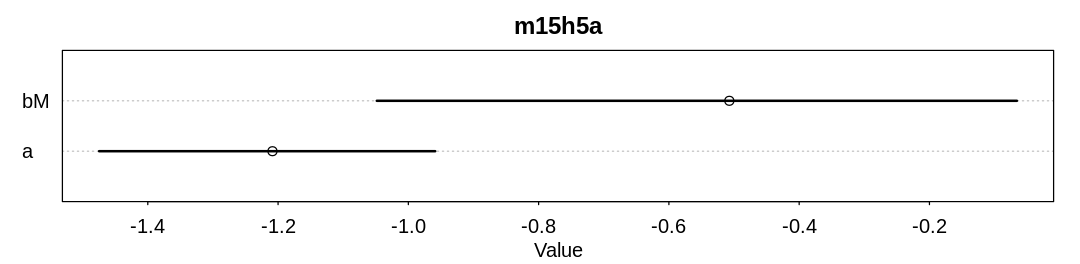
The negative intercept a merely indicates that most values are 0 rather than 1 in D (not missing).
More interesting is bM, which is reliably negative. A negative bM indicates that larger values
of M are associated with being missing less. That is, brain sizes are missing more often for
species with small masses.
Clearly this is not MCAR. See section 15.2.2 for potential DAGs. If you believe that brain volumes are only missing because researchers prefer to study species with a larger mass, then this may be a salvageable MAR situation. If you believe brain volumes are missing because researchers prefer to study species with larger brains, this is MNAR.
Found 56 NA values in B and attempting imputation.
Raw data (following plot):
mean sd 5.5% 94.5% n_eff
a 0.429209747 0.0570991572 0.339948276 0.520651112 889.8638
b 0.783647438 0.0138930038 0.762394288 0.805935970 948.3123
sigma 0.293430363 0.0156388062 0.269724497 0.318633929 1072.5169
B_impute[1] 0.004786200 0.0014914046 0.002857822 0.007525111 1137.7001
B_impute[2] 0.029086884 0.0083617833 0.018039909 0.043735841 1575.0531
B_impute[3] 0.034563101 0.0103652766 0.021077614 0.053294099 1017.0016
B_impute[4] 0.026491233 0.0082289481 0.015560821 0.040971634 1489.8989
B_impute[5] 0.225915045 0.0726967881 0.130827723 0.349635062 836.4177
B_impute[6] 0.032250991 0.0094624808 0.019980671 0.048312020 1516.9083
B_impute[7] 0.037880390 0.0118272347 0.022293129 0.058300099 1398.9808
B_impute[8] 0.034554255 0.0108554389 0.020622710 0.053117521 1520.0070
B_impute[9] 0.045598981 0.0131587578 0.027158449 0.068392413 1863.5973
B_impute[10] 0.041774107 0.0128845153 0.024896337 0.064855272 1125.7264
B_impute[11] 0.018121363 0.0054244604 0.010799199 0.027650251 2035.6780
B_impute[12] 0.014231789 0.0044881272 0.008344933 0.022448822 1586.1794
B_impute[13] 0.015329548 0.0046909879 0.009262177 0.023614649 1571.3961
B_impute[14] 0.016122983 0.0047035829 0.009709436 0.024484244 2297.1859
B_impute[15] 0.016484752 0.0049648826 0.009806028 0.025120559 1808.9191
B_impute[16] 0.018505499 0.0053459654 0.011731652 0.027859439 1797.5330
B_impute[17] 0.093398955 0.0280276883 0.055897370 0.143725021 1457.8011
B_impute[18] 0.130130424 0.0408690371 0.076146237 0.203106584 999.3712
B_impute[19] 0.129359063 0.0383762879 0.078740431 0.199986927 1201.2827
B_impute[20] 0.070147251 0.0211930144 0.042256754 0.106495169 866.6901
B_impute[21] 0.012061878 0.0037404611 0.007220431 0.018747894 1621.0253
B_impute[22] 0.050166179 0.0152995489 0.030274479 0.077493567 1254.9425
B_impute[23] 0.860432847 0.2620480707 0.513400500 1.325823939 1116.8501
B_impute[24] 0.142664219 0.0460889820 0.082237482 0.215528521 812.1062
B_impute[25] 0.025412435 0.0075556927 0.015203344 0.039167057 1689.2951
B_impute[26] 0.028134694 0.0081404623 0.017316218 0.042867656 1299.7887
B_impute[27] 0.169314478 0.0546392671 0.099653470 0.260259193 819.2582
B_impute[28] 0.093156644 0.0287961199 0.056129526 0.143993327 982.3735
B_impute[29] 0.116278165 0.0360459097 0.068648519 0.179953866 1137.0522
B_impute[30] 0.221431434 0.0632861315 0.138755290 0.328337691 1196.3263
B_impute[31] 0.216952919 0.0652025838 0.127738585 0.333300396 1212.2576
B_impute[32] 0.002483832 0.0007899518 0.001431282 0.003865538 1293.6024
B_impute[33] 0.004399372 0.0013400503 0.002625177 0.006772746 1195.9556
B_impute[34] 0.002301103 0.0006781731 0.001391275 0.003527719 1508.4254
B_impute[35] 0.003809495 0.0011719956 0.002227190 0.006039362 1356.6416
B_impute[36] 0.003345134 0.0010734592 0.001956075 0.005216269 1472.9661
B_impute[37] 0.004320865 0.0012847241 0.002571459 0.006529500 1509.1870
B_impute[38] 0.041955488 0.0121875359 0.025458276 0.063306769 1582.9235
B_impute[39] 0.152073431 0.0461289557 0.090522707 0.234132139 1215.9228
B_impute[40] 0.167228264 0.0503866096 0.100458543 0.253082815 1251.5582
B_impute[41] 0.017628984 0.0054501432 0.010653640 0.026924360 1646.1347
B_impute[42] 0.229739458 0.0675851325 0.139328718 0.349393552 1091.0886
B_impute[43] 0.193920914 0.0597911106 0.116504261 0.302735805 1090.9389
B_impute[44] 0.180186612 0.0512559379 0.109414988 0.272320041 1329.3292
B_impute[45] 0.068089177 0.0210701081 0.040722215 0.105434886 853.7422
B_impute[46] 0.094615481 0.0285466364 0.056778052 0.145485725 1245.0110
B_impute[47] 0.205792596 0.0675916585 0.119281179 0.324160656 803.9435
B_impute[48] 0.233534784 0.0740091353 0.136150060 0.369226377 1175.6142
B_impute[49] 0.252462732 0.0768465961 0.147657038 0.381118539 1226.3441
B_impute[50] 0.019421691 0.0058181517 0.011540242 0.029780839 2163.5233
B_impute[51] 0.017648020 0.0053207744 0.010650871 0.027172169 1294.9259
B_impute[52] 0.016768843 0.0050340439 0.009744324 0.025868531 2547.8019
B_impute[53] 0.029890798 0.0091762387 0.017554664 0.046480812 1613.0035
B_impute[54] 0.032199437 0.0098086945 0.018890907 0.049411995 1640.9744
B_impute[55] 0.209300633 0.0602691992 0.129023668 0.313635790 1205.6605
B_impute[56] 0.183497458 0.0561860323 0.109550257 0.283267329 976.8538
Rhat4
a 1.0012201
b 1.0007573
sigma 1.0025655
B_impute[1] 1.0003645
B_impute[2] 1.0007252
B_impute[3] 1.0054650
B_impute[4] 1.0000994
B_impute[5] 0.9991215
B_impute[6] 1.0013527
B_impute[7] 1.0005014
B_impute[8] 1.0022631
B_impute[9] 0.9998690
B_impute[10] 1.0006931
B_impute[11] 1.0001202
B_impute[12] 1.0011720
B_impute[13] 0.9994300
B_impute[14] 0.9999795
B_impute[15] 0.9991311
B_impute[16] 0.9996744
B_impute[17] 1.0008510
B_impute[18] 1.0034034
B_impute[19] 1.0015601
B_impute[20] 1.0010288
B_impute[21] 1.0001575
B_impute[22] 0.9998241
B_impute[23] 1.0017472
B_impute[24] 1.0001907
B_impute[25] 1.0026519
B_impute[26] 1.0000780
B_impute[27] 1.0001801
B_impute[28] 1.0011353
B_impute[29] 1.0003357
B_impute[30] 1.0053738
B_impute[31] 1.0007850
B_impute[32] 1.0033039
B_impute[33] 1.0031479
B_impute[34] 0.9997142
B_impute[35] 1.0052562
B_impute[36] 1.0018235
B_impute[37] 1.0016042
B_impute[38] 0.9997651
B_impute[39] 0.9999042
B_impute[40] 0.9998815
B_impute[41] 0.9989642
B_impute[42] 1.0030449
B_impute[43] 1.0011753
B_impute[44] 1.0005502
B_impute[45] 1.0019154
B_impute[46] 0.9996109
B_impute[47] 1.0002261
B_impute[48] 1.0019765
B_impute[49] 0.9993826
B_impute[50] 0.9998903
B_impute[51] 1.0008995
B_impute[52] 0.9989648
B_impute[53] 0.9990433
B_impute[54] 0.9997337
B_impute[55] 0.9995189
B_impute[56] 1.0000349
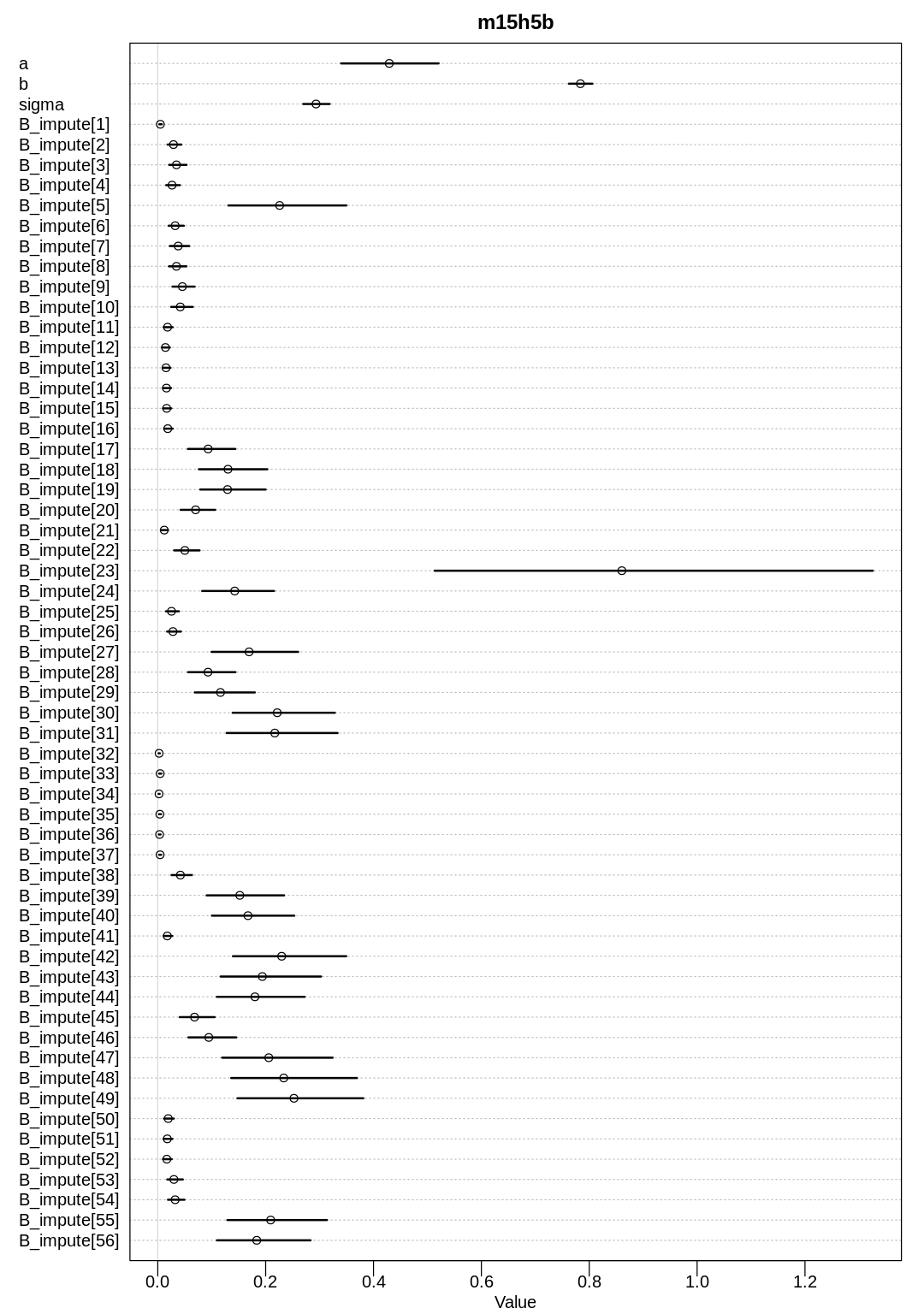

Running MCMC with 4 parallel chains, with 1 thread(s) per chain...
Chain 1 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 1 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 1 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 1 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 1 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 1 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 1 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 1 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 1 Exception: lognormal_lpdf: Scale parameter is inf, but must be positive finite! (in '/tmp/RtmpjPjDJw/model-1268e8b4e7.stan', line 18, column 4 to column 32)
Chain 1 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 1 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 1
Chain 1 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 1 Exception: lognormal_lpdf: Scale parameter is inf, but must be positive finite! (in '/tmp/RtmpjPjDJw/model-1268e8b4e7.stan', line 18, column 4 to column 32)
Chain 1 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 1 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 1
Chain 1 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 1 Exception: lognormal_lpdf: Scale parameter is inf, but must be positive finite! (in '/tmp/RtmpjPjDJw/model-1268e8b4e7.stan', line 18, column 4 to column 32)
Chain 1 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 1 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 1
Chain 1 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 1 Exception: lognormal_lpdf: Scale parameter is 0, but must be positive finite! (in '/tmp/RtmpjPjDJw/model-1268e8b4e7.stan', line 18, column 4 to column 32)
Chain 1 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 1 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 1
Chain 2 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 2 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 2 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 2 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 2 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 2 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 2 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 2 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 2 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 2 Exception: lognormal_lpdf: Scale parameter is 0, but must be positive finite! (in '/tmp/RtmpjPjDJw/model-1268e8b4e7.stan', line 18, column 4 to column 32)
Chain 2 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 2 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 2
Chain 3 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 3 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 3 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 3 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 3 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 3 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 3 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 3 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 3 Exception: lognormal_lpdf: Scale parameter is inf, but must be positive finite! (in '/tmp/RtmpjPjDJw/model-1268e8b4e7.stan', line 18, column 4 to column 32)
Chain 3 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 3 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 3
Chain 4 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 4 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 4 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 4 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 4 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 4 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 4 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 4 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 4 Exception: lognormal_lpdf: Scale parameter is 0, but must be positive finite! (in '/tmp/RtmpjPjDJw/model-1268e8b4e7.stan', line 18, column 4 to column 32)
Chain 4 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 4 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 4
Chain 1 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 1 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 1 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 1 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 1 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 1 finished in 0.3 seconds.
Chain 2 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 2 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 2 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 2 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 3 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 3 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 3 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 3 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 3 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 4 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 4 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 4 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 4 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 2 finished in 0.3 seconds.
Chain 3 finished in 0.3 seconds.
Chain 4 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 4 finished in 0.4 seconds.
All 4 chains finished successfully.
Mean chain execution time: 0.3 seconds.
Total execution time: 0.6 seconds.
Raw data (following plot):
mean sd 5.5% 94.5% n_eff Rhat4
a 0.4267492 0.05525369 0.3397942 0.5146442 923.0049 1.003886
b 0.7830870 0.01328885 0.7622681 0.8047012 899.6270 1.003091
sigma 0.2929341 0.01539626 0.2687894 0.3185917 705.9164 1.001046
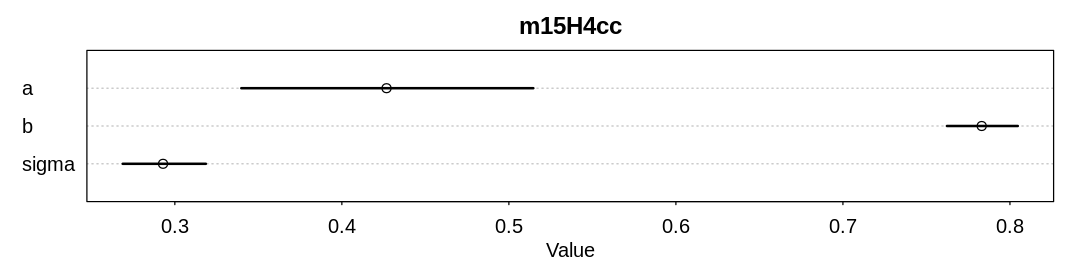
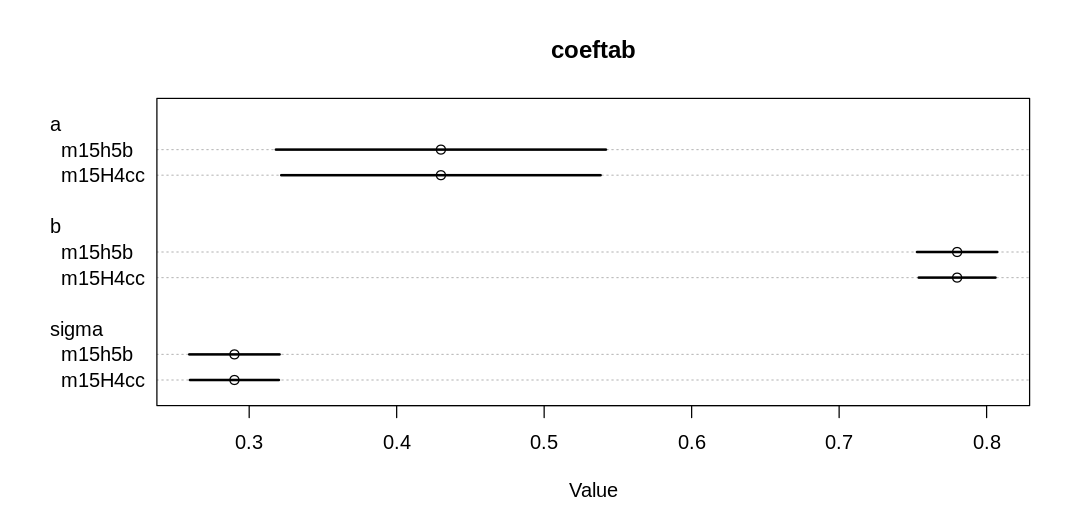
There doesn’t seem to be any change in the inferences for a, b, and sigma. When you impute
output values, the model isn’t really doing anything but posterior predictive inference. We haven’t
supplied any new assumptions about what the distribution of B should look like, other than what we
know from M (which we only learned from complete cases).
15H6. Return to the divorce rate measurement error model. This time try to incorporate the full generative system: \(A \rightarrow M \rightarrow D\), \(A \rightarrow D\). What this means is that the prior for \(M\) should include \(A\) somehow, because it is influenced by \(A\).
load.d15h6 <- function() {
data(WaffleDivorce)
d <- WaffleDivorce
dat <- list(
D_obs = standardize(d$Divorce),
D_sd = d$Divorce.SE / sd(d$Divorce),
M_obs = standardize(d$Marriage),
M_sd = d$Marriage.SE / sd(d$Marriage),
A = standardize(d$MedianAgeMarriage),
N = nrow(d)
)
return(dat)
}
a15h6a <- function(dat) {
m15.2 <- ulam(
alist(
D_obs ~ dnorm(D_true, D_sd),
vector[N]:D_true ~ dnorm(mu, sigma),
mu <- a + bA * A + bM * M_true[i],
M_obs ~ dnorm(M_true, M_sd),
vector[N]:M_true ~ dnorm(0, 1),
a ~ dnorm(0, 0.2),
bA ~ dnorm(0, 0.5),
bM ~ dnorm(0, 0.5),
sigma ~ dexp(1)
),
cmdstan = TRUE, data = dat, chains = 4, cores = 4
)
display_precis(m15.2, "m15.2", ar=0.4)
return(m15.2)
}
q15h6b <- function(dat) {
m15h6 <- ulam(
alist(
D_obs ~ dnorm(D_true, D_sd),
vector[N]:D_true ~ dnorm(mu, sigma),
mu <- a + bA * A + bM * M_true[i],
M_obs ~ dnorm(M_true, M_sd),
vector[N]:M_true ~ dnorm(mu_M, 1),
mu_M <- aM + bAM * A,
aM ~ dnorm(0, 0.2),
a ~ dnorm(0, 0.2),
bA ~ dnorm(0, 0.5),
bM ~ dnorm(0, 0.5),
bAM ~ dnorm(0, 0.5),
sigma ~ dexp(1)
),
cmdstan = TRUE, data = dat, chains = 4, cores = 4
)
display_precis(m15h6, "m15h6", ar=0.4)
return(m15h6)
}
e15h6b <- r"(
Notice `bAM` is nonzero. However, most of the original parameters are unchanged, as expected based
on the discussion in section **5.1.4** (`M` adds little predictive power once you know `A`).
)"
q15h6 <- function() {
dat <- load.d15h6()
display_markdown("**Answer.** Let's start by reproducing results from the chapter:")
m15.2 <- a15h6a(dat)
display_markdown("Fitting the new model:")
m15h6 <- q15h6b(dat)
iplot(function() {
plot(coeftab(m15.2, m15h6), pars=c("a", "bA", "bM", "sigma"), main="coeftab")
}, ar=2)
display_markdown(e15h6b)
}
q15h6()
Answer. Let’s start by reproducing results from the chapter:
Running MCMC with 4 parallel chains, with 1 thread(s) per chain...
Chain 1 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 1 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 1 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 1 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 2 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 2 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 2 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 2 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 2 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 2 Exception: normal_lpdf: Scale parameter is 0, but must be positive! (in '/tmp/RtmpjPjDJw/model-123eff970e.stan', line 28, column 4 to column 34)
Chain 2 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 2 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 2
Chain 3 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 3 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 3 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 3 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 4 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 4 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 4 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 4 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 4 Exception: normal_lpdf: Scale parameter is 0, but must be positive! (in '/tmp/RtmpjPjDJw/model-123eff970e.stan', line 28, column 4 to column 34)
Chain 4 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 4 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 4
Chain 1 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 1 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 1 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 1 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 1 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 2 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 2 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 2 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 2 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 2 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 2 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 2 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 3 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 3 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 3 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 3 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 3 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 4 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 4 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 4 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 4 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 4 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 1 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 1 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 1 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 2 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 3 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 3 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 3 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 4 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 4 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 1 finished in 0.4 seconds.
Chain 2 finished in 0.3 seconds.
Chain 3 finished in 0.4 seconds.
Chain 4 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 4 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 4 finished in 0.4 seconds.
All 4 chains finished successfully.
Mean chain execution time: 0.4 seconds.
Total execution time: 0.6 seconds.
Raw data (following plot):
mean sd 5.5% 94.5% n_eff Rhat4
D_true[1] 1.12385798 0.3554066 0.54481399 1.68852375 1914.401 0.9998953
D_true[2] 0.74050110 0.5266029 -0.08015318 1.59452610 2444.684 0.9998991
D_true[3] 0.42053533 0.3161150 -0.07594199 0.93404446 2597.937 0.9991635
D_true[4] 1.46071123 0.4480914 0.74302428 2.18795620 2510.655 0.9986221
D_true[5] -0.89449025 0.1259901 -1.09231825 -0.68827442 2127.438 0.9997797
D_true[6] 0.68512870 0.3867411 0.07769300 1.30872455 2292.443 0.9995023
D_true[7] -1.35476021 0.3430152 -1.90592495 -0.80999384 2481.670 0.9996085
D_true[8] -0.28698893 0.4548214 -1.01598670 0.41175814 2475.836 0.9990076
D_true[9] -1.78497443 0.6018430 -2.71190090 -0.82044548 1997.348 0.9997020
D_true[10] -0.62441650 0.1650684 -0.89387593 -0.36438402 2698.861 0.9987466
D_true[11] 0.77751538 0.2935488 0.31978693 1.25578840 2016.897 0.9987995
D_true[12] -0.45209473 0.4729781 -1.20371080 0.29501435 1803.887 1.0021392
D_true[13] 0.24294624 0.4715200 -0.52307459 0.96934553 1402.331 1.0011768
D_true[14] -0.86766586 0.2254476 -1.22821000 -0.49401419 2411.601 0.9997595
D_true[15] 0.54340814 0.3058431 0.05794533 1.03787930 2339.175 0.9983663
D_true[16] 0.28457517 0.3785022 -0.32169396 0.88385075 2560.544 1.0009084
D_true[17] 0.51067483 0.4163774 -0.15068532 1.16342025 3012.344 0.9995116
D_true[18] 1.23537672 0.3389570 0.70068147 1.78937960 2185.490 1.0013140
D_true[19] 0.43054590 0.3816789 -0.16673172 1.05539540 2935.632 0.9996269
D_true[20] 0.24361644 0.5390114 -0.60930384 1.11169005 1259.548 1.0034180
D_true[21] -0.55886426 0.3156862 -1.06725495 -0.04662007 2947.657 0.9994248
D_true[22] -1.10886942 0.2473762 -1.50504785 -0.71464697 2560.147 1.0022444
D_true[23] -0.30507948 0.2487208 -0.71469129 0.08518569 2185.574 1.0019136
D_true[24] -1.02378336 0.2922501 -1.48872680 -0.57042309 2238.040 0.9990170
D_true[25] 0.40349368 0.3942681 -0.20790610 1.02558770 1876.685 0.9994361
D_true[26] -0.04755081 0.3053162 -0.54331310 0.43034702 2843.097 0.9993076
D_true[27] -0.04985283 0.5001203 -0.85829017 0.74741533 2852.683 1.0006701
D_true[28] -0.14513949 0.4056516 -0.81104229 0.48514383 2881.513 0.9993679
D_true[29] -0.30315273 0.4723143 -1.03334420 0.47123007 1925.885 0.9998698
D_true[30] -1.80815678 0.2307198 -2.17006860 -1.43145570 2293.208 0.9984504
⋮ ⋮ ⋮ ⋮ ⋮ ⋮ ⋮
M_true[25] -0.1448684243 0.37293504 -0.74455669 0.43709568 1892.4066 0.9989351
M_true[26] -0.3811124354 0.20494118 -0.71277557 -0.04695257 2586.3208 0.9997921
M_true[27] -0.3235572561 0.52736772 -1.15066815 0.53674010 2911.4103 0.9982081
M_true[28] -0.1433853533 0.35678454 -0.71350194 0.44161150 2615.8537 0.9991915
M_true[29] -0.7045548847 0.40187257 -1.33865510 -0.03454238 2315.0763 0.9989793
M_true[30] -1.3776183825 0.15088344 -1.62501045 -1.13820930 2282.7380 0.9999063
M_true[31] 0.0660791122 0.43749275 -0.61072549 0.75275760 2219.3813 1.0007310
M_true[32] -0.8663235600 0.12741769 -1.06660815 -0.66400894 2411.1577 0.9998176
M_true[33] 0.0717017755 0.24752616 -0.32552679 0.45669929 3181.0730 0.9986132
M_true[34] 0.9689018099 0.62724536 -0.03917871 1.99773575 2405.8041 0.9990945
M_true[35] -0.8213976310 0.16609261 -1.09209135 -0.55897318 2767.5670 0.9994034
M_true[36] 0.9071350519 0.33482186 0.35913469 1.45461885 2784.3482 0.9983553
M_true[37] -0.2667911624 0.27913679 -0.71303866 0.16792863 2635.3455 1.0004072
M_true[38] -1.1997689575 0.12348601 -1.39374705 -1.00127745 3088.8690 0.9997407
M_true[39] -1.0074613184 0.46519207 -1.76555605 -0.27849233 2504.4498 1.0007309
M_true[40] -0.5011537417 0.29221312 -0.95379836 -0.02462128 2572.5901 1.0001853
M_true[41] -0.0007725454 0.57542887 -0.92589931 0.88127204 2416.7046 0.9987366
M_true[42] -0.1637778936 0.22125186 -0.51520496 0.18682741 2693.1536 0.9985345
M_true[43] 0.3513243068 0.15965578 0.10107341 0.60185929 2658.6843 0.9982000
M_true[44] 1.9525186970 0.43109855 1.29961745 2.64893650 2469.4158 0.9997747
M_true[45] -0.6865810001 0.52788819 -1.50918005 0.17976398 2595.0752 0.9986795
M_true[46] 0.0930301573 0.20905387 -0.24043334 0.43013554 2097.9095 0.9993468
M_true[47] 0.3251173919 0.24122606 -0.05922683 0.71370753 2507.4179 0.9990207
M_true[48] 0.4699208470 0.39142002 -0.15237856 1.09293820 2236.3509 1.0014293
M_true[49] -0.7466404448 0.19900366 -1.06804200 -0.43654851 2829.3651 0.9995158
M_true[50] 1.2858845407 0.70408722 0.17539639 2.45705750 2117.5642 1.0017119
a -0.0425067809 0.09662695 -0.19308193 0.11207957 1432.8261 0.9984967
bA -0.5483560592 0.16292837 -0.81169573 -0.28882952 1081.4427 1.0017287
bM 0.2020118926 0.21267107 -0.12967567 0.53664554 758.0593 1.0051126
sigma 0.5510849735 0.10375394 0.38863759 0.72575790 619.3232 1.0077815
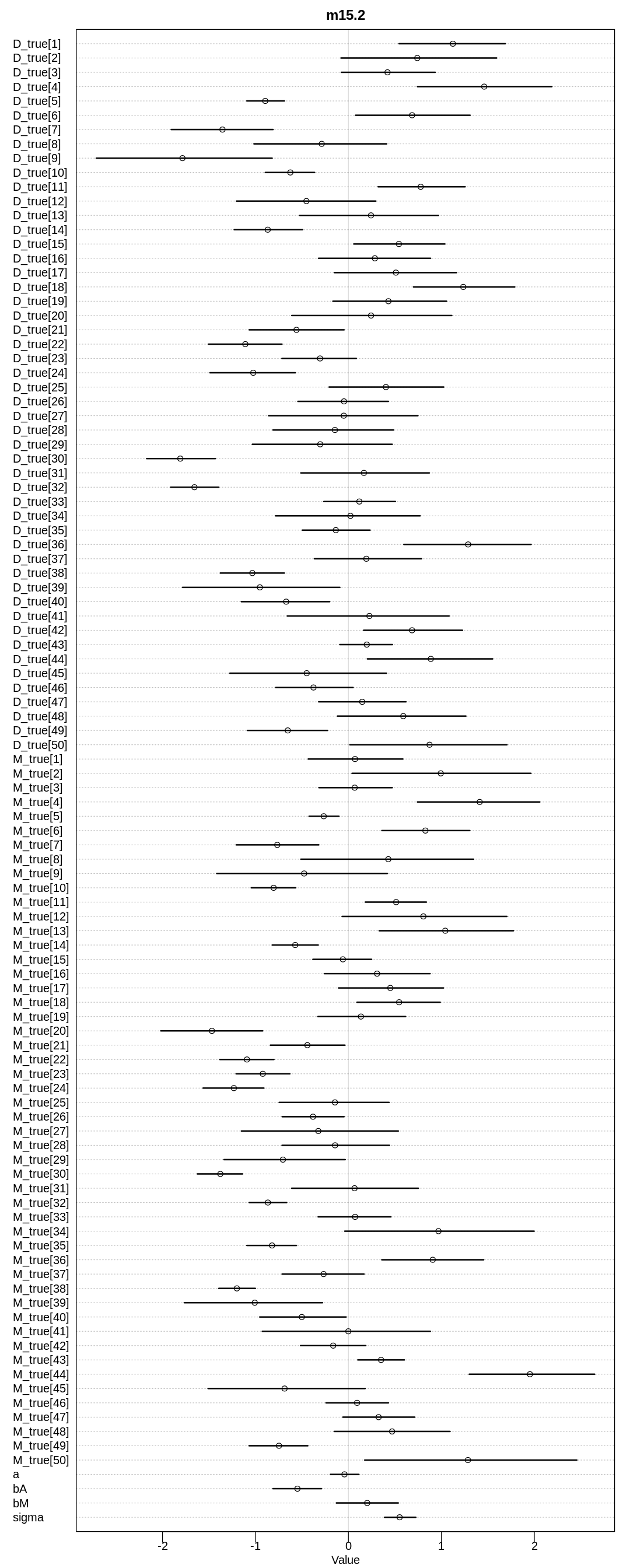
Fitting the new model:
Running MCMC with 4 parallel chains, with 1 thread(s) per chain...
Chain 1 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 1 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 1 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 2 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 2 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 2 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 2 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 2 Exception: normal_lpdf: Scale parameter is 0, but must be positive! (in '/tmp/RtmpjPjDJw/model-12249236b8.stan', line 36, column 4 to column 34)
Chain 2 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 2 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 2
Chain 3 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 3 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 3 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 4 Iteration: 1 / 1000 [ 0%] (Warmup)
Chain 4 Iteration: 100 / 1000 [ 10%] (Warmup)
Chain 1 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 1 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 1 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 1 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 1 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 2 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 2 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 2 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 2 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 2 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 3 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 3 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 3 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 4 Iteration: 200 / 1000 [ 20%] (Warmup)
Chain 4 Iteration: 300 / 1000 [ 30%] (Warmup)
Chain 4 Iteration: 400 / 1000 [ 40%] (Warmup)
Chain 1 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 1 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 2 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 2 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 2 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 2 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 3 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 3 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 3 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 3 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 4 Iteration: 500 / 1000 [ 50%] (Warmup)
Chain 4 Iteration: 501 / 1000 [ 50%] (Sampling)
Chain 4 Iteration: 600 / 1000 [ 60%] (Sampling)
Chain 2 finished in 0.4 seconds.
Chain 1 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 1 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 3 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 3 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 4 Iteration: 700 / 1000 [ 70%] (Sampling)
Chain 4 Iteration: 800 / 1000 [ 80%] (Sampling)
Chain 4 Iteration: 900 / 1000 [ 90%] (Sampling)
Chain 1 finished in 0.5 seconds.
Chain 3 finished in 0.4 seconds.
Chain 4 Iteration: 1000 / 1000 [100%] (Sampling)
Chain 4 finished in 0.5 seconds.
All 4 chains finished successfully.
Mean chain execution time: 0.5 seconds.
Total execution time: 0.7 seconds.
Raw data (following plot):
mean sd 5.5% 94.5% n_eff Rhat4
D_true[1] 1.15741184 0.3595291 0.59412252 1.74972665 2074.339 1.0006261
D_true[2] 0.73003959 0.5469587 -0.12951053 1.60639965 2306.957 0.9987878
D_true[3] 0.43048803 0.3482953 -0.11874783 0.99947320 2745.864 0.9996444
D_true[4] 1.45810499 0.4762043 0.70619835 2.21900915 2612.755 0.9990083
D_true[5] -0.89844902 0.1289282 -1.10140970 -0.69021863 2891.040 0.9991381
D_true[6] 0.68066719 0.3973069 0.06299353 1.30106485 3260.033 0.9987280
D_true[7] -1.36457105 0.3540565 -1.94241340 -0.80054167 2338.352 0.9995367
D_true[8] -0.32822943 0.4806232 -1.12201045 0.41759172 1921.289 1.0005649
D_true[9] -1.86883291 0.6021902 -2.81836470 -0.88254134 1784.200 1.0021782
D_true[10] -0.62529896 0.1579800 -0.87109117 -0.37367789 3233.745 0.9991621
D_true[11] 0.76868559 0.2796956 0.33666835 1.22805045 1956.473 0.9994012
D_true[12] -0.52527207 0.4899950 -1.29960710 0.24368810 2053.003 1.0006026
D_true[13] 0.16622776 0.4815465 -0.61262437 0.90119968 1303.675 0.9997353
D_true[14] -0.86828884 0.2237228 -1.23626130 -0.50700680 2547.783 0.9999712
D_true[15] 0.53343476 0.2985346 0.06041430 1.02060690 2433.651 0.9997813
D_true[16] 0.28778024 0.3777882 -0.31387357 0.88401106 2227.041 0.9993436
D_true[17] 0.51062931 0.4274562 -0.13550758 1.19631340 2896.135 0.9992179
D_true[18] 1.25345616 0.3455893 0.71119767 1.81143085 2220.506 0.9995943
D_true[19] 0.43898114 0.3791973 -0.15190421 1.04743770 2311.989 0.9985908
D_true[20] 0.35956008 0.5607253 -0.47115521 1.29438130 1256.289 1.0016227
D_true[21] -0.55297449 0.3135544 -1.06210115 -0.04361986 2774.016 1.0009602
D_true[22] -1.09450045 0.2523780 -1.48913400 -0.69456875 2808.485 0.9988513
D_true[23] -0.28616501 0.2611433 -0.71201247 0.13880295 2067.197 0.9988679
D_true[24] -1.00610083 0.2906452 -1.47546390 -0.54541531 1919.609 0.9992381
D_true[25] 0.42796329 0.4121669 -0.23009722 1.09007990 2678.018 1.0002640
D_true[26] -0.04345925 0.3177689 -0.54807182 0.43438509 2494.603 1.0001580
D_true[27] -0.03687699 0.4940858 -0.84317314 0.75180598 2415.242 0.9996449
D_true[28] -0.16204792 0.3966675 -0.79008256 0.48044281 2991.713 0.9985323
D_true[29] -0.27487396 0.5107981 -1.05462010 0.56081215 2084.902 1.0010248
D_true[30] -1.81449135 0.2447264 -2.20513550 -1.42679180 2331.857 0.9990249
⋮ ⋮ ⋮ ⋮ ⋮ ⋮ ⋮
M_true[27] -0.28923060 0.54690892 -1.18023410 0.58945606 2698.8430 0.9987006
M_true[28] -0.09182479 0.35302763 -0.65251846 0.45166202 2785.4455 0.9987070
M_true[29] -0.80201348 0.41152929 -1.44995700 -0.14778865 2602.0373 0.9989405
M_true[30] -1.39175353 0.15530592 -1.63198830 -1.14317020 2697.6732 0.9998723
M_true[31] 0.08217810 0.42530037 -0.58524281 0.76108709 2623.9126 1.0007984
M_true[32] -0.87636522 0.12236965 -1.06934135 -0.68060106 2646.8222 0.9995971
M_true[33] 0.07919226 0.25055348 -0.33089932 0.48249644 2726.8911 0.9987757
M_true[34] 1.15444743 0.60967329 0.15492086 2.12917970 3131.7459 0.9990234
M_true[35] -0.83005297 0.15572244 -1.07748365 -0.58163307 3003.4485 0.9986447
M_true[36] 0.97891253 0.32159342 0.43837574 1.49501505 2601.8884 1.0001305
M_true[37] -0.28830185 0.27470838 -0.73073531 0.15900299 3000.0338 0.9985948
M_true[38] -1.20582236 0.12291540 -1.39910685 -1.00237735 3103.4038 0.9984800
M_true[39] -1.27723121 0.47164140 -2.01509910 -0.52995943 2484.1396 0.9983936
M_true[40] -0.51197200 0.29538098 -0.99158171 -0.04300853 3171.2424 1.0000859
M_true[41] 0.06659838 0.58581477 -0.86348428 1.00052695 2868.5150 0.9995583
M_true[42] -0.14762759 0.21755571 -0.49363252 0.18497697 2665.3211 1.0004558
M_true[43] 0.36461604 0.15730744 0.11083406 0.61445859 2989.6685 0.9998724
M_true[44] 2.23985825 0.43964666 1.54402570 2.93478195 2075.8127 0.9995402
M_true[45] -0.82197830 0.56628402 -1.75761095 0.08632466 2600.6192 0.9986284
M_true[46] 0.07905651 0.21048304 -0.25326136 0.40616174 2929.1503 0.9989778
M_true[47] 0.32459071 0.24795602 -0.07911612 0.72100663 3165.4918 0.9992154
M_true[48] 0.55095543 0.38791439 -0.08567644 1.15856975 2700.9058 1.0004629
M_true[49] -0.75362239 0.20906057 -1.07695540 -0.41069346 2919.1697 0.9992028
M_true[50] 1.79173403 0.71098989 0.65998150 2.93193095 2709.0857 0.9992667
aM -0.03838471 0.12467626 -0.23042831 0.16585141 2454.9284 0.9996420
a -0.04708609 0.09969729 -0.20777371 0.10764634 1872.9229 0.9986087
bA -0.56984227 0.17168467 -0.83871434 -0.29010205 868.6785 1.0064443
bM 0.12587028 0.20792856 -0.19685901 0.46149217 696.3589 1.0062410
bAM -0.63835680 0.14670222 -0.87692010 -0.40137013 1877.6531 0.9994998
sigma 0.58195392 0.11223721 0.41170270 0.76920186 613.5693 1.0022078
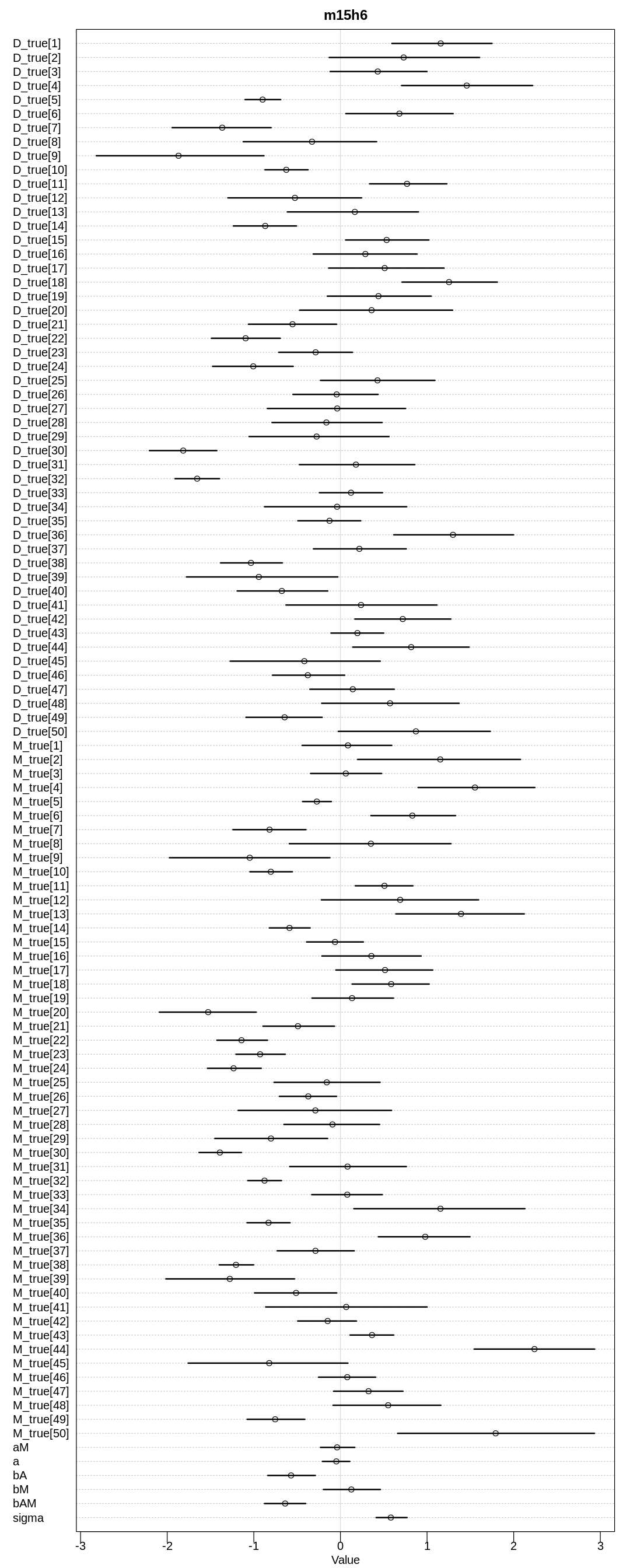
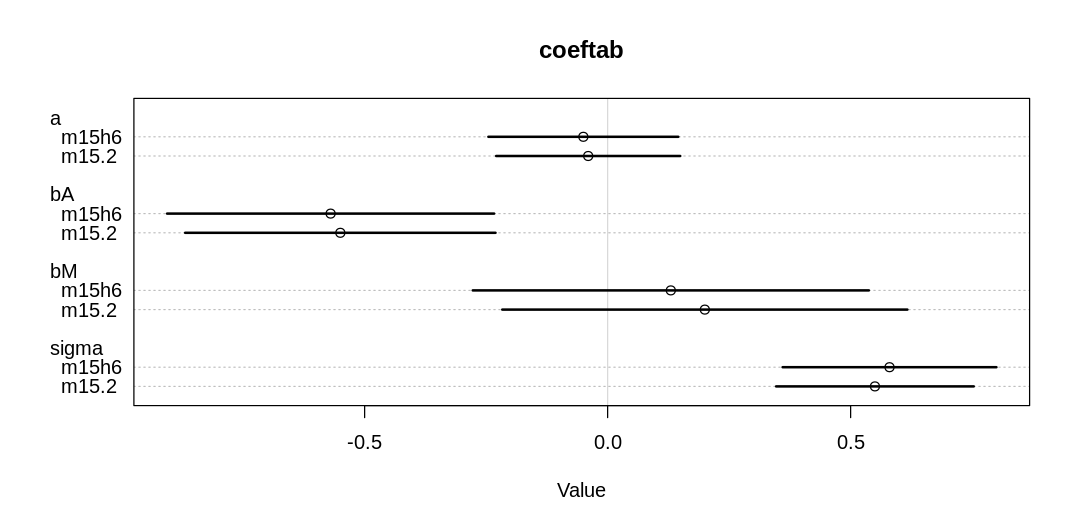
Notice bAM is nonzero. However, most of the original parameters are unchanged, as expected based
on the discussion in section 5.1.4 (M adds little predictive power once you know A).
15H7. Some lad named Andrew made an eight-sided spinner. He wanted to know if it is fair. So he spun it a bunch of times, recording the counts of each value. Then he accidentally spilled coffee over the 4s and 5s. The surviving data are summarized below.
library(data.table)
load.d15h7 <- function() {
V = seq(8)
Freq = c(18, 19, 22, NA, NA, 19, 20, 22)
d <- data.frame(Value = V, Frequency = Freq)
t_d <- transpose(d)
colnames(t_d) <- rownames(d)
rownames(t_d) <- colnames(d)
display(t_d)
return(d)
}
e15h7b <- r"(
Your job is to impute the two missing values in the table above. Andrew doesn’t remember how many
times he spun the spinner. So you will have to assign a prior distribution for the total number of
spins and then marginalize over the unknown total. Andrew is not sure the spinner is fair (every
value is equally likely), but he’s confident that none of the values is twice as likely as any
other. Use a Dirichlet distribution to capture this prior belief. Plot the joint posterior
distribution of 4s and 5s.
)"
e15h7c <- r"(
**Answer.** We could model this as a categorical distribution where we predict the category i.e. the
spinner value from no data; we don't need a predictor to spin. To do this, we'd have to expand the
given table into many trials. How would we model a variable number of observations, though?
To model a variable number of observations, it seems like we'd need to use the multinomial
distribution. It doesn't look like this distribution is available through `ulam`. It's also not
clear how we'd deal with what is essentially a single observation in this case.
What if you took the $Value$ as a categorical predictor? The assumption that no value is twice as
likely as any other would become the variance on a prior. But the Dirichlet distribution strongly
implies either a categorical or multinomial.
To plot the joint posterior distribution, use `pairs` on the model.
Skipping this question for now.
)"
q15h7 <- function() {
d <- load.d15h7()
display_markdown(e15h7b)
display_markdown(e15h7c)
}
q15h7()
| 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | |
|---|---|---|---|---|---|---|---|---|
| <dbl> | <dbl> | <dbl> | <dbl> | <dbl> | <dbl> | <dbl> | <dbl> | |
| Value | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 |
| Frequency | 18 | 19 | 22 | NA | NA | 19 | 20 | 22 |
Your job is to impute the two missing values in the table above. Andrew doesn’t remember how many times he spun the spinner. So you will have to assign a prior distribution for the total number of spins and then marginalize over the unknown total. Andrew is not sure the spinner is fair (every value is equally likely), but he’s confident that none of the values is twice as likely as any other. Use a Dirichlet distribution to capture this prior belief. Plot the joint posterior distribution of 4s and 5s.
Answer. We could model this as a categorical distribution where we predict the category i.e. the spinner value from no data; we don’t need a predictor to spin. To do this, we’d have to expand the given table into many trials. How would we model a variable number of observations, though?
To model a variable number of observations, it seems like we’d need to use the multinomial
distribution. It doesn’t look like this distribution is available through ulam. It’s also not
clear how we’d deal with what is essentially a single observation in this case.
What if you took the $Value$ as a categorical predictor? The assumption that no value is twice as likely as any other would become the variance on a prior. But the Dirichlet distribution strongly implies either a categorical or multinomial.
To plot the joint posterior distribution, use pairs on the model.
Skipping this question for now.